Awesome 
I saw today that v1 of the fastai library was available as Kaggle Kernels.
I think Kernels are a great learning platform–earlier this year I adapted the fast.ai Part 1 v2 notebooks to run in the Kernels environment: https://towardsdatascience.com/announcing-fast-ai-part-1-now-available-as-kaggle-kernels-8ef4ca3b9ce6.
I got the lesson1-pets notebook from v3 running as a Kernel, and you can find it here: https://www.kaggle.com/hortonhearsafoo/fast-ai-v3-lesson-1. Really the only “trick” that was required this time around was to set num_workers=0 for the Databunches, as there is an issue with Pytorch(?), seen in other platforms too, that breaks in multi-worker situations.
Great work! Congratulations!
I would like to ask you if the inference was done on mobile or in the cloud?
Thanks!
Thanks. I guess I need to run more experiments 
I created another starter pack  This time it is for the Kaggle Humpback Whale Identification Competition. I like it way more than the previous one. Also, the competition is a lot of fun - many things could be attempted that can be a good learning experience and that can lead to a better score. The competition just launched and there are still over two months and a half to go!
This time it is for the Kaggle Humpback Whale Identification Competition. I like it way more than the previous one. Also, the competition is a lot of fun - many things could be attempted that can be a good learning experience and that can lead to a better score. The competition just launched and there are still over two months and a half to go!
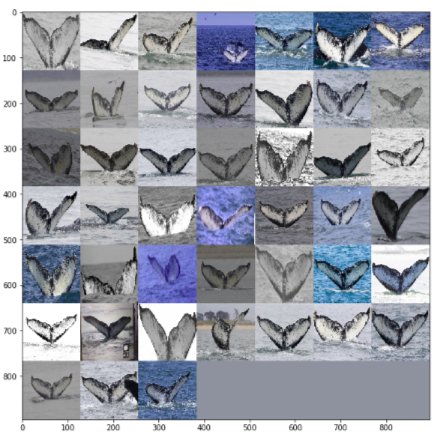
Here are a couple of images from the training set:

Sorry for not being able to share an awesome project like everyone else.
Instead I want to share an interview: I had interviewed @rachel about her DL Journey and fast.ai, I totally forgot to share it with the community. Here is the link to the interview.
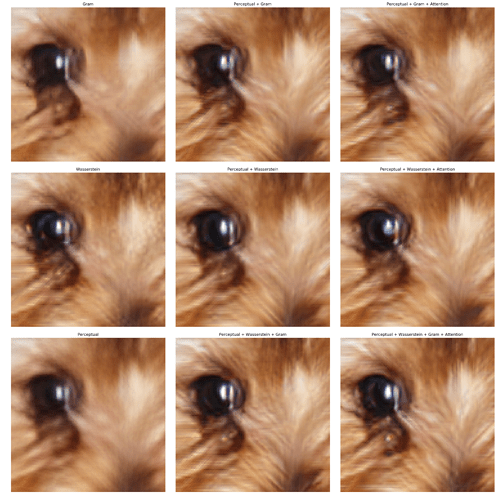
Based on the crap to no crap GAN of lesson 7, I tried the same approach to add colors to a crap black & white image. First I downloaded high quality images from EyeEm using this approach.
Training the Generator with simple MSE loss function, I got not exciting results (the only thing it learned is sky has to be BLUE  ):
):
Then after adding the Discriminator I and training in a ping-pong fashion, the Generator got better:
Now trying on some test images 
I replied on Kaggle, but wanted to reply here as well: Your methodology very closely followed the intuitions I’ve been working off of, and your code gave me a couple of "ah hah!"s from my own experiments so far. You can see my detailed response there.
Thanks for posting the useable map5 code also.
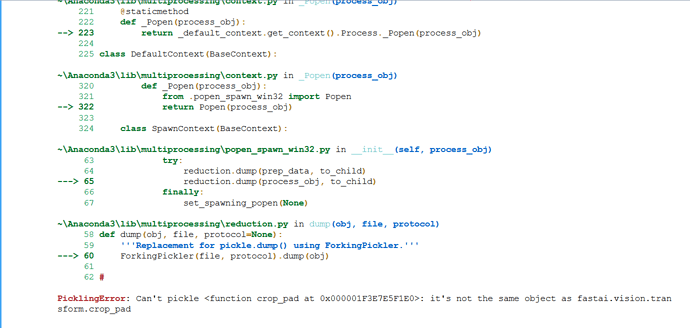
@radek. Thank you for sharing the notebook. I am trying to run the notebook. I got the error
PicklingError: Can’t pickle <function crop_pad at 0x000001F3E7E5F1E0>: it’s not the same object as fastai.vision.transform.crop_pad
I am using FastAi library 1.0.36.post1 on windows 10.
Please suggest how to fix this error.
Thanks,
Ritika Agarwal
Chances are just restarting the kernel should fix this.
I answered here - for what works for me at least with Windows 10
Worked perfectly- thanks for sharing this!
@radek I have restarted the kernel.Then also I am getting this error. I fixed this issue by putting NUM_WORKERS =0 and adding padding_mode=‘zeros’
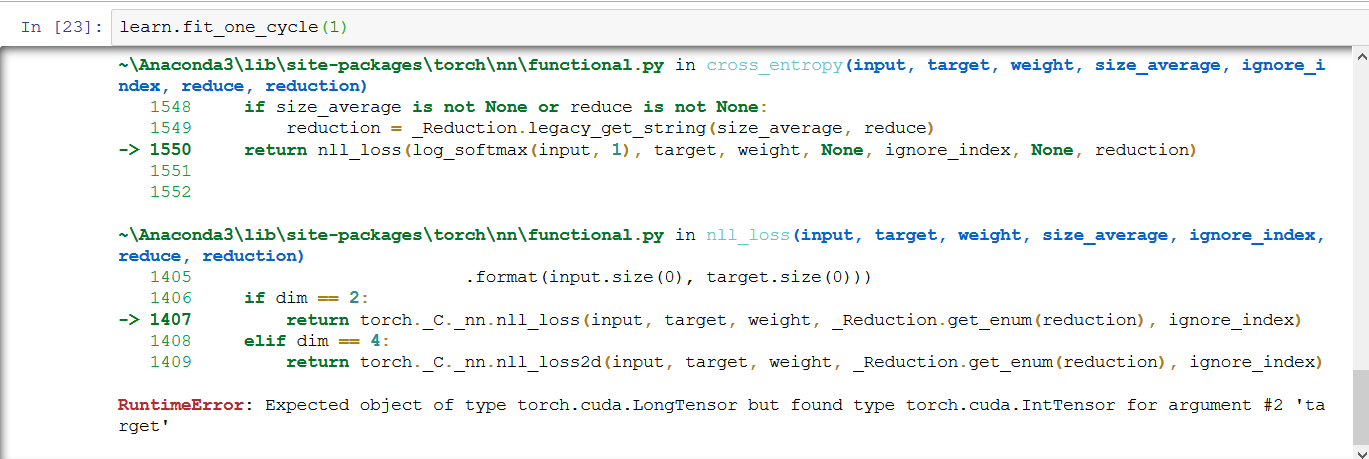
But Now i am getting the below error while fitting the model. Pls suggest how to fix this error.
RuntimeError: Expected object of type torch.cuda.LongTensor but found type torch.cuda.IntTensor for argument #2 ‘target’
Thanks,
Riitka
Thanks @brismith. I have replicated the steps shared by you. Still I got this error.
I fixed this issue by adding NUM_WORKERS =0 and adding padding_mode=‘zeros’
Thanks,
Ritika
I played around with different loss functions for super resolution to see the impact on the generated images. Interesting to see that impacts quality. I’m also trying to figure out how to deploy a super resolution model on Zeit. If anyone knows how to make the app return an image, let me know.
That means your fastai version is not up to date.
I  a brief post on the Naive Bayes classifier (Introduced in the fastai Machine Learning Course, Lesson #10) on Towards Data Science. Hoping some folks might find it useful.
a brief post on the Naive Bayes classifier (Introduced in the fastai Machine Learning Course, Lesson #10) on Towards Data Science. Hoping some folks might find it useful.
Hi Jeremy,
I am using fastai version 1.0.36.post1 on windows 10. Do i need to update fastai package?
Thanks,
Ritika
One way to do this is export the .ipynb in Jupiter as a .py script.
Then you can browse (and jump to source) or execute the script in vscode.