Hi mates!
I am struggling with unexpected (for me) behaviour of fast.ai library.
I am trying to reuse Lesson 3 code for another dataset.
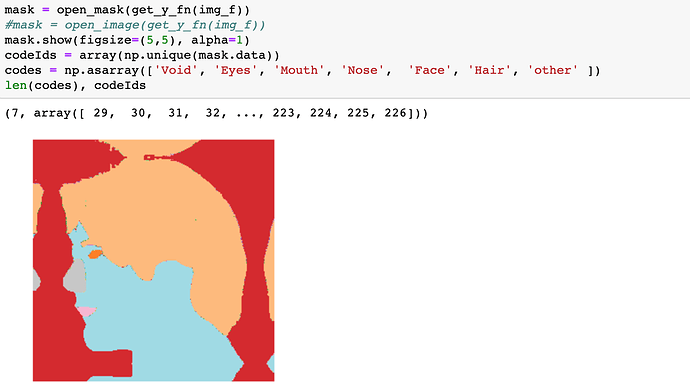
Initially dataset has a lot of classes per mask:
We need only 7 classes (face, nose, eyes …)
So I tried to cleanup label files with preprocessing to assign label values from 0 to 6,
having 0 for background like this:
classes = [76, 37, 225, 178, 149, 29, 58] # original classes
#76 => background
#37 => hair
def cls_cleanup(inp):
file = np.array(inp)
for key, cls in enumerate(classes):
m = file == cls
file[m]= key
file[file > len(classes)] = 0
return Image.fromarray(file)
....
for fld in inp.ls():
f = Path(fld)
o = out/f.name
if not os.path.exists(o):
os.makedirs(o)
for file in f.ls():
if 'labels' not in str(file.parent):
s_f = open_image(file)
else:
s_f = open_mask(file, after_open=cls_cleanup)
o_f = crop_pad(s_f, size, 'reflection', 0.5,0.5)
o_f.save(o/file.name.replace('.bmp', '.png'))
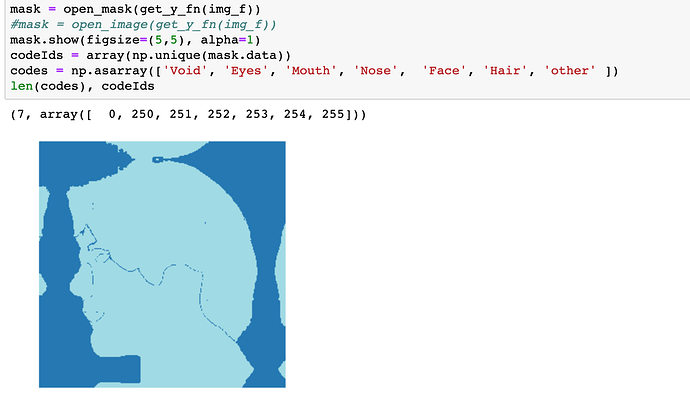
after such preprocessing, newly loaded masks somehow
have labels: 0, 250,251,252,253,254,255
but we need 0,1,2,3,4,5,6
Your help will be really appreciated, I am fighting with this issue almost a week

//Viktor
P.S. Looks like such class indexes are causing such error once trying to get learning rate:
---------------------------------------------------------------------------
RuntimeError Traceback (most recent call last)
~/.local/lib/python3.6/site-packages/fastai/basic_train.py in fit(epochs, learn, callbacks, metrics)
99 xb, yb = cb_handler.on_batch_begin(xb, yb)
--> 100 loss = loss_batch(learn.model, xb, yb, learn.loss_func, learn.opt, cb_handler)
101 if cb_handler.on_batch_end(loss): break
~/.local/lib/python3.6/site-packages/fastai/basic_train.py in loss_batch(model, xb, yb, loss_func, opt, cb_handler)
31 if opt is not None:
---> 32 loss,skip_bwd = cb_handler.on_backward_begin(loss)
33 if not skip_bwd: loss.backward()
~/.local/lib/python3.6/site-packages/fastai/callback.py in on_backward_begin(self, loss)
288 "Handle gradient calculation on `loss`."
--> 289 self.smoothener.add_value(loss.detach().cpu())
290 self.state_dict['last_loss'], self.state_dict['smooth_loss'] = loss, self.smoothener.smooth
RuntimeError: CUDA error: device-side assert triggered