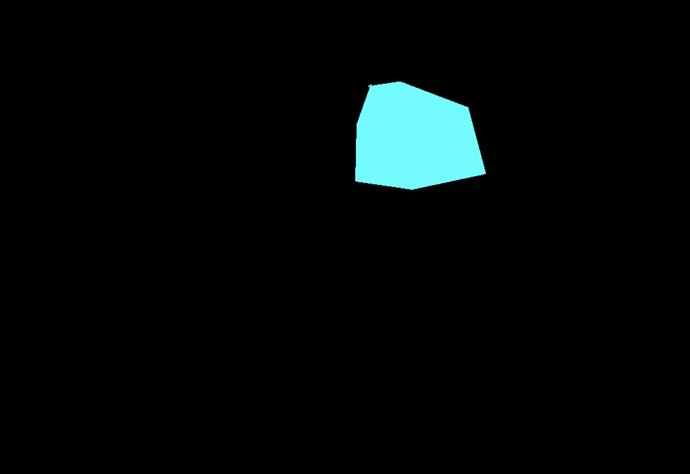
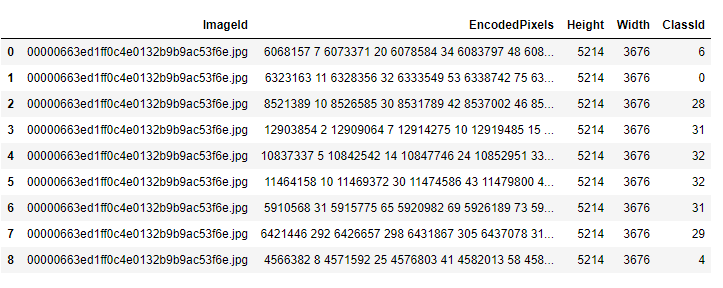
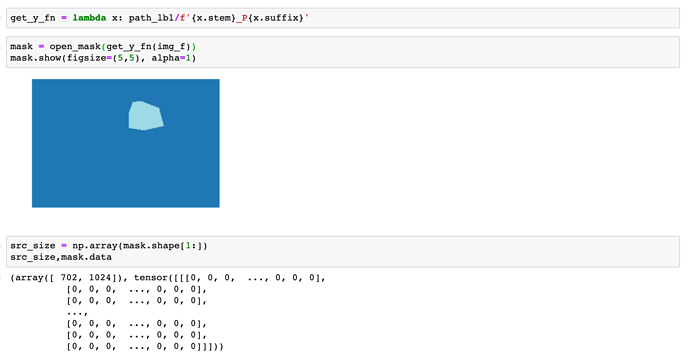
Many seem to have problems with Segmentation and the used masks. Because of that I wrote some functions that can help anybody to overcome the problems. I did not test it thoroughly, only one RGB and one gray scaled image, where it worked as expected.
You only have to provide the path, where the labels / masks are as well as a path to a folder where you want to save the converted masks. The new files will be named as the old ones. Converted masks contain only values from 0 to N, where N is the number of classes -1.
Here the code:
%reload_ext autoreload
%autoreload 2
%matplotlib inline
from fastai.vision import *
from fastai.callbacks.hooks import *
import PIL.Image as PilImage
def getClassValues(label_names):
containedValues = set([])
for i in range(len(label_names)):
tmp = open_mask(label_names[i])
tmp = tmp.data.numpy().flatten()
tmp = set(tmp)
containedValues = containedValues.union(tmp)
return list(containedValues)
def replaceMaskValuesFromZeroToN(mask,
containedValues):
numberOfClasses = len(containedValues)
newMask = np.zeros(mask.shape)
for i in range(numberOfClasses):
newMask[mask == containedValues[i]] = i
return newMask
def convertMaskToPilAndSave(mask,
saveTo):
imageSize = mask.squeeze().shape
im = PilImage.new('L',(imageSize[1],imageSize[0]))
im.putdata(mask.astype('uint8').ravel())
im.save(saveTo)
def convertMasksToGrayscaleZeroToN(pathToLabels,
saveToPath):
label_names = get_image_files(pathToLabels)
containedValues = getClassValues(label_names)
for currentFile in label_names:
currentMask = open_mask(currentFile).data.numpy()
convertedMask = replaceMaskValuesFromZeroToN(currentMask, containedValues)
convertMaskToPilAndSave(convertedMask, saveToPath/f'{currentFile.name}')
print('Conversion finished!')
Now you only have to use:
convertMasksToGrayscaleZeroToN(pathToLabels, saveToPath)
I did not tune the code for high performance, it should just do what hopefully helps a lot of people here.
Any suggestions for adding / changing parts are welcome.