Hi,
I am trying to run the lesson notebook but when I execute the import I get an error “No module names ‘bcolz’” which is already installed in my system!
Error in notebook!
Checking installation of Bcolz
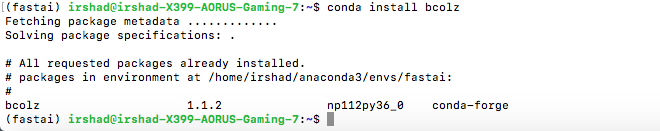
Need help to solve the issue.
Hi,
I am trying to run the lesson notebook but when I execute the import I get an error “No module names ‘bcolz’” which is already installed in my system!
Error in notebook!

Need help to solve the issue.
Hmm, one thing you might try is to double check that the jupyter command you ran to launch the notebook is installed in your fastai virtualenv/conda environment, versus being some other installation of jupyter. That is, double check that running which jupyter at the command line prints something like some/path/to/envs/fastai/bin/jupyter, as opposed to something global like ~/anaconda/bin/jupyter. You could also run pip freeze from within your virtualenv and see if jupyter is in there.
I’ve made that mistake before!
Thanks man, it worked like a charm 
Cheers
Hi Irshad can you please elaborate what worked for you? I’m getting the same error, when I did pip freeze it shows me jupyter is installed.
What is confusing to me is that Jeremy’s script does a “source activate fastai” but I don’t see it creating a conda environment called “fastai” … when I’m within this so called fastai env, and do which jupyter, it points to the global install not local install…
Nevermind, the reason I didn’t have anything installed in the fastai conda environment was because I ran the script manually line by line on my local box and I think I may have skipped the part where I need to be in the fastai directory to run the ‘conda env update’ command.
I issued the “conda env update” command from within the already activated fastai env that the script somehow creates. It installed bcolz as part of installing all the packages etc.
Once it was done I deactivated and reactivated the fastai env and jupyter notebook was able to find bcolz.
cheers!
I have tried a bunch of solutions listed below but still get the same error. Any ideas?
conda install -c anaconda bcolz
pip install conda
source activate my_conda_env
(my_conda_env) $ conda install nb_conda
ModuleNotFoundError Traceback (most recent call last)
in ()
1 # This file contains all the main external libs we’ll use
----> 2 from fastai.imports import *
~/sample-notebooks/fastai/courses/dl1/fastai/imports.py in ()
1 from IPython.lib.deepreload import reload as dreload
----> 2 import PIL, os, numpy as np, math, collections, threading, json, bcolz, random, scipy, cv2
3 import random, pandas as pd, pickle, sys, itertools, string, sys, re, datetime, time, shutil
4 import seaborn as sns, matplotlib
5 import IPython, graphviz, sklearn_pandas, sklearn, warnings
ModuleNotFoundError: No module named ‘bcolz’
You should update your envirorment.
Go to your working directory and type:
conda env update
And then activate:
conda activate fastai
found this thread while resolving same problem.
my blog below documents this and other fixes to get the python notebook running.
I’ve documented a series of setup issues on this medium account. Hope these help a future someone 
Hii,
I have found the same error while doing setup even when I had already installed the required packages, after checking forums again I tried to run the jupyter notebook as @cqfd suggested from the working directory then I was not getting this issue anymore.
Thanks a lot @cqfd
Thanks, it worked !!!
If this is the issue, how do i correct the path if i do not have sudo permissions?
I went ahead and install Jupyter Notebook in my fastai cuda env and it worked 
I changed directory to my fastai root, activated conda env, and install jupyter notebook there
cd ~/fastai
source activate fastai
conda install jupyter
conda env update
source deactivate
source activate fastai
jupyter notebookare you able to solve this error?
ah awesome. this finally worked. I’m a prorammer well versed in bash, but couldn’t figure this thing out…Feel sorry for the less experienced people 
Thanks. This worked for my Windows 10.
I got this same error. A dumb and quick thing that I tried and worked was changing the ipykernel to the default (Python 3).
Hi guys, I would really appreciate some help here. I believe I have tried every suggestion here, but I still get the same error.
My system:
I’m sure I have this bcolz library installed. I can load it using ipython:
(fastai) C:\Users\gusta\source\repos\fastai>ipython
Python 3.6.8 |Anaconda, Inc.| (default, Dec 30 2018, 18:50:55) [MSC v.1915 64 bit (AMD64)]
Type 'copyright', 'credits' or 'license' for more information
IPython 7.2.0 -- An enhanced Interactive Python. Type '?' for help.
In [1]: import bcolz
In [2]:
I can even open the Jupyter notebook, and check that the package is visible there:
(Notice from the figure that the package list was obtained from the very shell that is running lesson1.ipynb. Jupyter is installed in the fastai environment. Still, when trying to load the cell, I get the same error:
I’m out of ideas here. Could anyone help me? I really appreciate any leads.
Thank you very much!
doesnot work for me… Any help.
have you find the solutions? I have the same problem with you too
on Mac Book (Pro) there in no support for gpu, so when installing you need to use ‘conda env create -f environment-cpu.yml’. that worked for me.