Towards the 1:30 min mark, I see Jeremy do in1 = F.relu(self.l_in(self.e(c1))). I thought that when we used embeddings we had to use the nn.ModuleList function. Why don’t we use it in this case?
Towards the 1:20 minute mark, I notice Jeremy talk about how the activation function gives more activations. I thought they gave the new weights. Is there a difference between activation and weight?
Also, I noticed Jeremy did this:
it = iter(md.trn_dl)
*xs,yt = next(it)
t = m(*V(xs))
How do you do m(*V(xs)) without fitting the model first? Am I missing something?
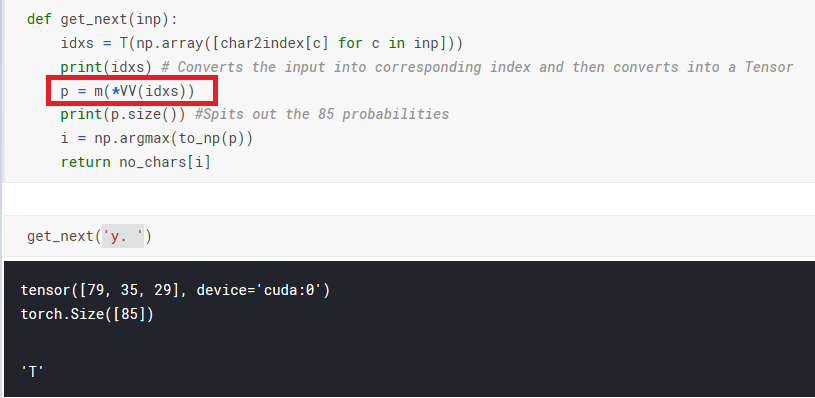
I ran into this. idxs needs to be a list of tensors so it can be broken up by * and fed into m(.).
def get_next(inp):
idxs = [T(np.array([char_indices[c]])) for c in inp]
p = m(*idxs)
i = np.argmax(to_np(p))
return chars[i]
Otherwise the first line of CharLoopConcatModel.forward is going to try to find the size of the first character index value (your entire n-length input string will be converted to a single n-length 1-D tensor).
Did you ever get an answer to your question since i have exactly the same issue, removing dim=-1 gives lower losses
Sorry, I never got around to figuring it out.
thanks for getting back
Does the Hinton paper referenced in the lesson (https://arxiv.org/abs/1504.00941) mean that we can now get results comparable to LSTM when using vanilla RNNs?
I think there is something that I don’t understand about the multi-output model and first characters of each batch.
In the video, Jermy (2:07:09) says that we just need to reset the hidden state to 0 when we initiate the model. The only character in the dataset that does not have the previous datapoint is the first character in the text.
But I changed the code and replaced the text with vector containing number representing the character order in the text( 1,2,3,4…128) and also changed batchsize to 4 and cs to 3 to make things more easy to follow.
When running next(iter(md.trn_dl)) the first time, i get the values:
itteration 1
x1 =[72 42 87 69]
x2 =[73 43 87 70]
x3 = [74 44 88 71]
itteration 2
x1 =[ 78 36 75 69]
x2 = [79 37 76 70]
x3 = [80 38 77 71]
But I expected the itteration 2 to return
x1 = [75 45 90 72]
x2 = [76 46 91 73]
x3 = [77 47 92 74]
The dataset is shuffled. But even if I set “shuffle=False” it makes no sense to me.
I thought that the columns of the data (eg forst char of x1,x2 and x3) to be continous between the batches to be able to use the hidden state in the model correctly.
Is this a bug or is it something that I need to understand better?
if I wanna extract the embedding matrix and use it for prediction in forest or decision tree, what’s my steps here?
I am thinking to do this, but I am not sure if this is correct
- Save the embedding matrix
- Convert it into a numpy array
- Extract the categorical value from the matrix
like in the shape of embedding matrix(year, 4)
we extract the corresponding year of vector of size(1,4) out - put the vector into a list and put it into a column pandas dataframe as a new feature (no. of sample, no. of feature)
- convert the dataframe into matrix and put it into sklearn.
so essentially we are replacing the categorical feature by the vector of that feature in the embedding matrix, right? Thanks a lot.
Hello Friends,
I was working on the Three Character model where given three input our model should predict the third output.
I’m having some confusion in the output part .
May I know what does *VV does here . I tried *V , I was getting the output . I didn’t get the purpose of two times Variable(V) over there.
Can I get some help here.
Thanks
hi there,
I am now trying line by line of the SGD section. I find something funny but I can’t explain given my limited knowledge of pytorch so far.
a = V(np.random.randn(1), requires_grad=True)
b = V(np.random.randn(1), requires_grad=True)
So then I tried to add a .cuda() after a and b (i.e. a = a.cuda() and b.cuda()), then the whole online gradient batch descent does not work anymore
learning_rate = 1e-3
for t in range(10000):
# Forward pass: compute predicted y using operations on Variables
loss = mse_loss(a,b,x,y)
if t % 1000 == 0: print(loss.data[0])
# Computes the gradient of loss with respect to all Variables with requires_grad=True.
# After this call a.grad and b.grad will be Variables holding the gradient
# of the loss with respect to a and b respectively
loss.backward()
# Update a and b using gradient descent; a.data and b.data are Tensors,
# a.grad and b.grad are Variables and a.grad.data and b.grad.data are Tensors
a.data -= learning_rate * a.grad.data
b.data -= learning_rate * b.grad.data
# Zero the gradients
a.grad.data.zero_()
b.grad.data.zero_()
it fails with a.grad.data is None type:
AttributeError: ‘NoneType’ object has no attribute ‘data’
I really try that out for fun, but I think that would help me to understand pytorch more. Could anyway shed some lights on it?
Any luck ?
I’m trying to get a better intuitive feel for embeddings and how backprop helps compute meaningful weights in the embeddings. We’ve talked about two tasks to create embedding vectors, 1) a neural net to compute embeddings directly, as part of a dot product of two embedding vectors, and 2), categorical variables that are represented as embeddings that are inputs in a task to compute a sales prediction, as an example. My questions:
For 1), I can see how embeddings are computed. The network computes a loss on the label each time and backprops to fix the weights of the embeddings to get the dot products of the embeddings closer to the label.
I’m confused on 2). In 2), the objective function for example might be to predict sales. But then how does backprop work to set meaningful weights in the categorical variable embedding vectors? This feels a bit indirect of a task to get the right weights, and much harder to interpret.
I just spent some time working through this lesson.
I have one lingering question.
Here’s the model from the notebook:
# Here's the model exactly copied from fastbook.
class LMModel3(Module):
def __init__(self, vocab_sz, n_hidden=64):
# Take note of this: the embedding dimensions are vocab_sz by n_hidden.
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.h_h = nn.Linear(n_hidden, n_hidden)
self.h_o = nn.Linear(n_hidden,vocab_sz)
self.h = 0
My question is: What do you do if you want to use a number for the embedding dimension that is not the same as the number of hidden neurons?
If you just try to use a unique number of embedding dimensions, you get RuntimeError: mat1 and mat2 shapes cannot be multiplied.
I expect the answer is to just use another linear layer to transform the output of your embedding layer into the shape that you need. But I wanted to ask if there was something else/better.
Here’s a notebook with my notes.