In the tabular lesson assignments, Jeremy talks about using NN Embeddings in the RF model. How exactly is that done? Just concatenate the embeddings and the continuous variables and input that into the RF?
Doesn’t that seem a little weird to do from an RF perspective?
I would recommend reading this notebook by @Pak https://github.com/Pak911/fastai-shared-notebooks/blob/master/interpret_tabular.ipynb he shows an example
Woah that’s a lot of code lol, here I was thinking it was going to be two lines of indexing and Concat. Thanks!
Trying here instead as advised by muellerzr…
I’m confused on the process of how categorical embeddings from a NN can be concatenated with continuous variables and used in a Random Forest. Specifically embedding shape (rows) do not fit that of the original input (e.g. learn.model.embeds.parameters() . , thus I’m unclear how these can be concatenated to form the X for a new model such as a RandomForest.
I have seen the lengthy notebook linked (section “RF vs NN”) though it’s not clear to me how it’s approaching this problem as it’s not sufficiently commented for a novice. If anyone who has already stepped through this code - or has the familiarity to read it directly - can shed some light with a few pointers on the problem, their benevolence would be much appreciated!
I plan on looking into this in the next few weeks with Walk with fastai, so if no one has posted a good explanation til then know that is in the works of my to do list 
Thanks muellerzr, much appreciated.
please write a message here when you done. It will be like a notification.
Many thanks!
I would also be interested in this!
Got the answer to this finally, we can use hooks:
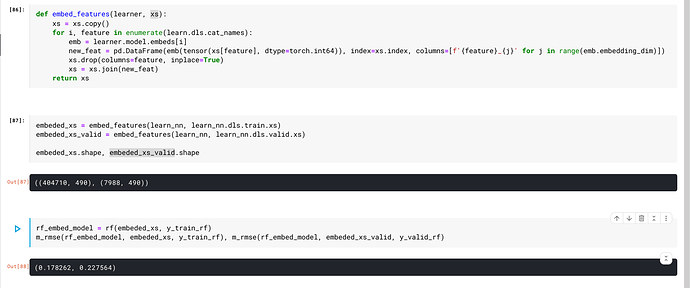
Here is what I do to use embeddings from a neural net in a random forest:
def embed_features(learner, xs):
xs = xs.copy()
for i, feature in enumerate(learn.dls.cat_names):
emb = learner.model.embeds[i]
new_feat = pd.DataFrame(emb(tensor(xs[feature], dtype=torch.int64)), index=xs.index, columns=[f'{feature}_{j}' for j in range(emb.embedding_dim)])
xs.drop(columns=feature, inplace=True)
xs = xs.join(new_feat)
return xs
I used that function then like this:
embeded_xs = embed_features(learn, learn.dls.train.xs)
xs_valid = embed_features(learn, learn.dls.valid.xs)
Hope this helps anyone!
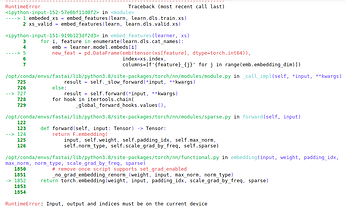
Hi, thank you so much for the solution. I have tried it but got:
RuntimeError: Input, output and indices must be on the current device.
I am stuck. I have checked stackoverflow for it but nothing worked out with torch.device("cuda" if torch.cuda.is_available() else "cpu") for me.
I am using gradient’s free GPUs.
Did you work with CPU? or maybe something wrong with gradient.
Has anyone tried nn embeddings on a rf and seen the model significantly improving?
I have tried but mse with and without embedding on the rf model is roughly the same.
Roughly getting (0.191887, 0.236464) as training mse, val mse for both.
@danielwbn
Thank you  Your function helped me to use embeddings in RF.
Your function helped me to use embeddings in RF.
@tsm_tau
I got my best RF result with RF with NN Embeddings version, althought it’s a small difference. The Neural Net model gave the best result.
Here are my results with all versions from the chapter in fastbook:
-
rf_model
Trial 1: (0.171001, 0.23253)
Trial 2: (0.171121, 0.232491) -
rf_model_imp
(0.181014, 0.230941)
(0.180963, 0.229838) -
rf_model_final
(0.183347, 0.232544)
(0.18334, 0.232704) -
rf_model_final_time
(0.191957, 0.228525)
(0.192147, 0.229181) -
rf_model_final_time_filt
(0.177479, 0.228915)
(0.177702, 0.22913) -
Neural Net
(None, 0.226417) -
rf_embed_model
(0.178262, 0.227564)
I got the same error when I trained Neural Net with GPU but not when I trained on CPU.
I think the problem is, when trained on GPU, the embeddings are still on GPU but when you try to use it on CPU, it throws the error that the data are not on same device.
It can be solved with a simple fix:
emb = learner.model.embeds[i].cpu()
Just move the data to CPU before using it!
this is great! thank you.
Thanks for sharing, but the link maybe be broken, is it only me?