I am here at room 153, Data Institute, but nobody is here. Am I in the correct room/place? 
Here now 
Couldn’t make it to SF today. But I hope to come in one day a week to hangout and possibly help.
Would anyone be interested in doing a coffee shop study group for maybe an hour in the morning? (I work full-time)
~8am?
If there’s any interest, we could talk about which days of the week!
I’m interested @jsonm ! Is the area near class convenient for you?
@brockinit I work near Montgomery Bart, so that’d be easy for me, but let’s see if anyone else is interested and try to pick a place that works for everyone!
@jsonm I’m definitely interested in study group!
Also, anyone interested in study together on weekend? I’m thinking about Saturday afternoon, Sunday afternoon or evening. Hit me up if you are interested!
Sophia
@sophia.onion @jsonm I’m definitely interested in a Weekend Study Group. I am based in the South Bay (San Jose) but could make the trek up on weekends. Weeknights can be a bit hard.
I am in San Francisco Monday, however, I work 9-5ish. Could cut out early and do a mid day or just before class. Thanks !
@sophia.onion @jsonm @fmichaelkunz I would be interested in a weekend study group. I’m also up for an evening study group (can’t do mornings). We could get together at a coffee shop or whatever.
I will be in San Jose for a few weekends during the course. Would love to join
I feel the same here. Why don’t we start doing a few mini PRs, get feedback, and find the best way to improve it (kind of like SGD itself lol)? Since the package and its dependencies gets updated so quickly, I’d start with documenting the stuff that’s unlikely to change in the next little while.
Hi, I need a help in a project. I have an idea to read ECG diagrams.
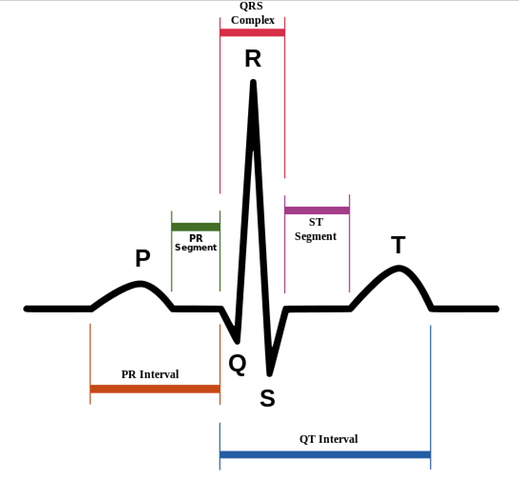
See this diagram.
ECG can be divided in to different waves, marked with different letters P, Q, R, S, and T, and this cycle repeats.
Assume, I have enough data (which I dont have in reality)
From a 30 seconds ECG, I want to separate each PQRST cycle, and in each block I want to mark the points P, Q, R, S and T.
Some time waves can be noise - this should be ignored
Some time, some of the components (PQRST) can be missing due to different illness conditions - this should be ignored.
So, my questions are,
- Is this really a deep learning problem?
- Which format of input is more efficient? As a time series or as Image?
- Can we use a bounding box to separate each cycle? and then use this bounding box to crop the image, and then find the PQRST points as X,Y co-ordinates in cropped sections running it through another neural network?
Can anybody help in this project to find a suitable architecture?
Refer : https://stanfordmlgroup.github.io/projects/ecg/
(Note: I am not a cardiologist, so my knowledge in ECG is very limited)
For those in SF looking for a weekend study group, note that there’s a separate thread for that now: http://forums.fast.ai/t/weekend-study-group-in-sf/13671
The SF study group yesterday successfully completed this terrific PR. Great to see such quick progress!
I am definitely interested Jason, I’ll be in SF since Tuesday, let me know!
Lots of successful work last week in the SF study group, including:
- The start of a documentation project
- New research in to architectures for segmentation
- Debugging memory leaks in fastai
FYI I won’t be in until late morning today, since I’m on toddler duty 
Looking forward to coming to SF tomorrow. I am hoping to try out what we learnt this week on a new dataset. Anyone has a dataset suggestion?
There’s a thread here on that exact topic - maybe a search for ‘coco’ will find it.
Awesome. Lots of great work - http://forums.fast.ai/t/other-datasets-to-predict-bounding-boxes-on/13795
Just added this google doc. Feel free to add other SF study group projects if you wish.