So here’s a draft todo list:
1. Exploratory Data Analysis (EDA)
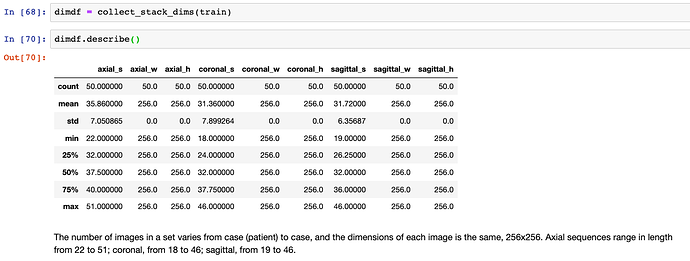
[x] Examine data format. For each case, three numpy array files (.npy), where each array comes from a scan along a different plane (axial, coronal, sagittal), and is of shape (s, 256, 256), corresponding to s images, or slices, each of dimension 256x256. The number of slices per plane differs from patient to patient.
[x] Visually examine examples.
[x] Note the probable scan techniques used for the data. Axial, coronal, and sagittal images use distinct methods.
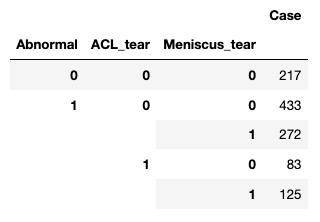
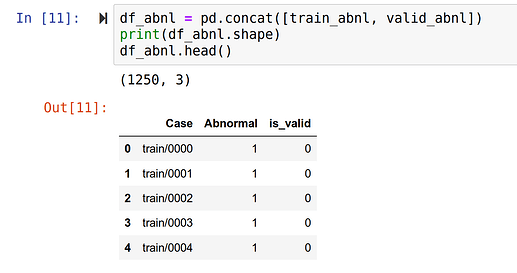
[x] Tabulate occurrence of various categories (normal, abnormal, ACL tear, Meniscus tear), noting class imbalances.
[ ] other EDA steps…
2. Prep data for modeling
[x] Convert files to images files for toy model.
[ ] Create custom ItemBase subclass.
[ ] Create custom ItemList subclass.
[ ] Create labeled DataBunch
[ ] Make sure data splitting for train/valid/test is appropriate (original paper authors kept all three scans per patient in same set.
[ ] Normalize data.
[ ] Explore adding appropriate data augmentation strategies (rotations, shifts, flips, perspective warping, superresolution, etc) possibly dependent on model used.
[ ] Other data prep steps…
3. Fit models to data
[ ] Start with toy model fit on middle slice from one plane, or from all three planes, using pre-trained 2D model.
[ ] Choose and implement loss function (original paper uses cross-entropy loss, re-scaled to account for class imbalances).
[ ] Implement competition’s target performance metric of AUC averaged across three classification tasks (detection of abnormality, of ACL tears, of Meniscal tears).
[ ] Replicate results of original paper using their model and data pre-processing approaches.
[ ] Improve on original paper by using fastai pre-processing and model tuning procedures.
[ ] Explore alternative model architectures for images and image sequences, where corresponding pre-trained model weights can be used.
[ ] Explore incorporating a segmentation step (ANT-GAN or other).
[ ] Explore volumetric, rather than 2d approaches.
[ ] Explore parameter and hyperparameter search to improve expected performance on out of sample data.
[ ] Other model fitting steps…
4. Submit results
[ ] Make official submission.
Lots to do. What do you want to work on?
 , so I’ve decided to use a Medium post as a sort of “lab notebook” for my data exploration (with included domain knowledge).
, so I’ve decided to use a Medium post as a sort of “lab notebook” for my data exploration (with included domain knowledge).