I’m happy to help. First, a little domain knowledge…
Here’s the description of the data (emphasis added):
Examinations were performed with GE scanners (GE Discovery, GE Healthcare, Waukesha, WI) with standard knee MRI coil and a routine non-contrast knee MRI protocol that included the following sequences: coronal T1 weighted, coronal T2 with fat saturation, sagittal proton density (PD) weighted, sagittal T2 with fat saturation, and axial PD weighted with fat saturation. A total of 775 (56.6%) examinations used a 3.0-T magnetic field; the remaining used a 1.5-T magnetic field.
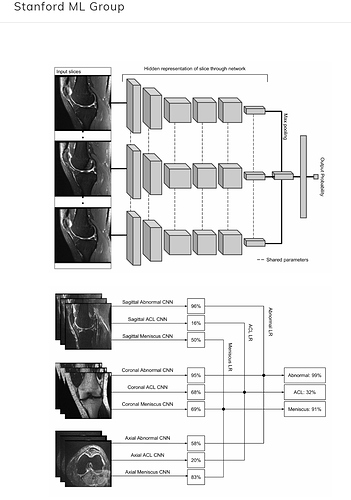
And here are the tasks:
The leaderboard reports the average AUC of the abnormality detection, ACL tear, and Meniscal tear tasks.
In a T1-weighted sequence, fluid will be intermediate-to-dark signal and fat will be bright. Fibrous structures, such as the ACL and meniscus - will be dark on all sequences. T2 and PD with fat saturation are referred to as fluid-sensitive sequences, because fluid stands out as bright signal on these images. Fat is normally bright on T2 and PD, but “fat saturation” saturates that signal - i.e. makes it dark - which makes it easier to see the fluid.
Abnormal fluid is often associated with pathology. There is normally some fluid in the knee joint capsule, but too much fluid often indicates a problem. Inflammation often results in increased fluid in the fat - called fat “stranding” because it looks wispy and strand-like. Menisci and ligaments (like the ACL) should be nice and dark. A focus of bright signal in one of these structures may indicate a tear.
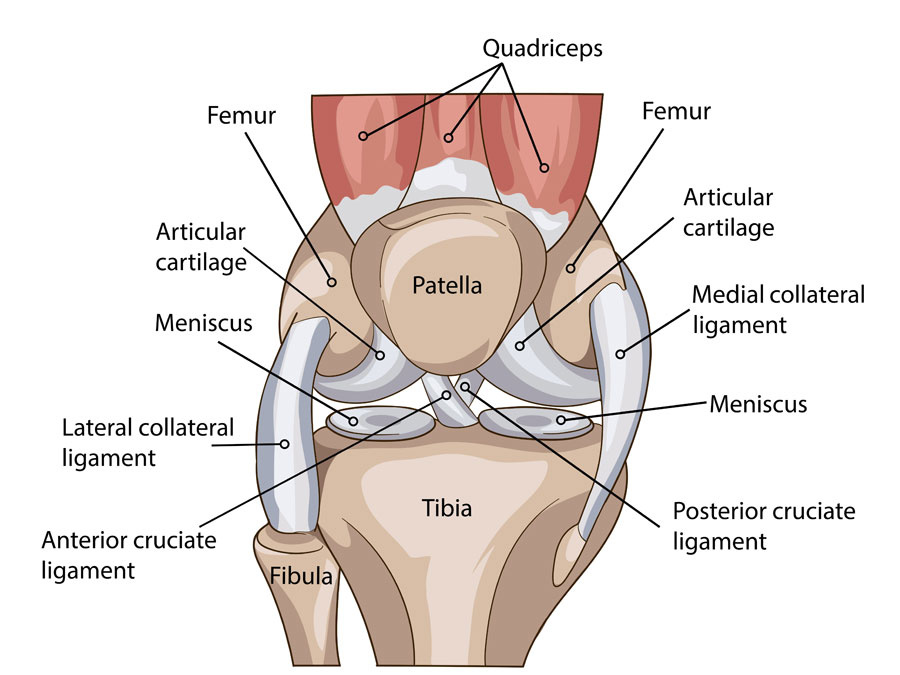
Super-basic knee anatomy:
I would suggest the following sequences for overall abnormality detection: coronal T1, sagittal PD and coronal T2 with fat saturation. The coronal T1 and sagittal PD will give you a nice overview of the anatomy, albeit with slightly lower sensitivity for pathology. The coronal T2 fat-sat provides a good overview screen for pathology due to the relative brightness of fluid.
The highest yield sequences for specifically detecting meniscal and ACL tears will most likely be the coronal T2 with fat-sat, sagittal T2 with fat-sat, and axial PD with fat-sat. Menisci and the ACL are typically better evaluated on the coronal and sagittal sequences, but the axial can provide useful information in certain edge cases. So a 3-channel model architecture might include each of these sequences as one input channel.
 That would be great bragging rights lol.
That would be great bragging rights lol.