Very good insights. Thanks for sharing
Was torch.tensor out of bounds or why did we use fastai’s tensor command instead of going directly to torch. Is there a benefit of using that command? Here is the line I’m talking about:
x_train,y_train,x_valid,y_valid = map(tensor, (x_train,y_train,x_valid,y_valid))
I would have instead used :
map(torch.tensor, (x_train,y_train,x_valid,y_valid))
(and let’s be honest I would have done each of those individually, who knew that map was a thing?)
Looked into the fastai tensor command a bit more and it looks like it actually uses torch.tensor so I’m guessing that is ok to use.
Is there a way to see the code of a built-in function or method?
I was going to look at what exactly torch.randn does to generate its random numbers with a normal distribution, but ??torch.randn doesn’t seem to work. It gives me the documentation but not the code.
torch.randn??
So the takeaway would be whenever you’re making a tensor of n similar iterables or applying a function to n iterables, just replace that with 1 single line of map(YourFunc, n1, n2....n)
Actually if you check the import section:
from torch import tensor
Which is why we can type tensor instead of torch.tensor
For other functions, if you look at the online PyTorch docs, there is an option that shows us the source code, but for torch.randn the same isn’t there either.
I’m not sure why it behaves this way, but for ?tensor and ??tensor and same for the randn function, both just return the docstring.
Thoughts on Lesson 8
This post summaries my ideas after the lesson and centres around these
elements.
- Normalisation
- Bayesian Methods
- Augmentation
Normalisation
The idea of normalising the data has significant weight in this lesson.
What we have done is normalise continuous variables, what would a
normalised categorical variable look like, would it be the same or not?.
Would this approach be readily applied to other learning strategies e.g.
- RandomForest Regression/Classifier
- GradientBoosting
- RandomTreeEmbedding
If we subject our data to mean of 0 and stdev of 1 then Bayesian methods
can be applied?.
Bayesian Methods
The figure below is taken from [^1] And it is a figure of a hierarchical
model taken from Chapter 3 Modeling with Linear Regression it shows a
series of distributions to make up a regression y = \alpha + X \beta
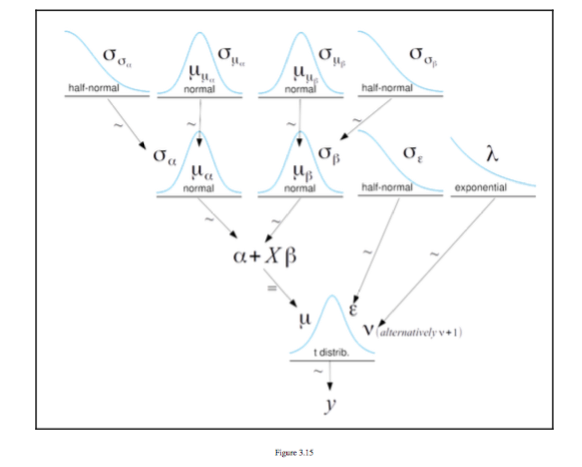
Augmentation
Given that we can create regressions from distributions and there are discrete
distributions for categories see [^1] could this be used for augmentation i some
way
I appreciate I may not have explained this very well but I am sure some discussion will help.
[^1]: Bayesian Analysis with Python Second Edition Author Osvaldo Martin
[^1]
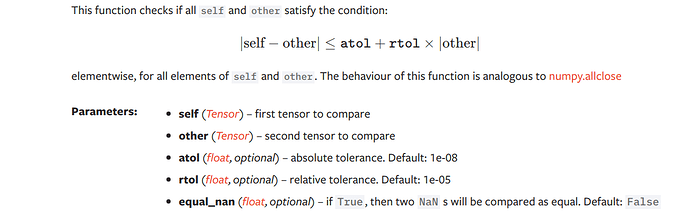
Any idea on what rtol and atol means in the below code
torch.allclose(a,b,rtol=1e-3,atol=1e-5)
From the docs:
atol (float, optional) – absolute tolerance. Default: 1e-08
rtol (float, optional) – relative tolerance. Default: 1e-05
Edit: I couldn’t make the equation render correctly, here is the screenshot of the equation:
It’s to correct for the mean of Relu not being zero. So its effect it to push the mean toward zero; in doing so, it also pushes the std towards 1. The latter happens because std depends on mean, so making the mean more accurate makes std more accurate as well.
Don’t forget you can use ‘??’ either before or after e.g. ‘torch.allclose??’ in a notebook cell and the method docs will appear. What my next question might be is what are atol or rtol controlling.
That was actually my question, I looked at the documentation before posting. I was able to understand it better after looking at the numpy documentation.
It looks to me from Sanyam’s post that the absolute difference between self and other has to be less than or equal to the absolute tolerance plus the relative tolerance times the absolute value of other.
This is where an intuition can be got by varying the parameters passed to the method which is what is part of Jeremy’s advice to us as learners print out the inputs/outputs and all that.
I hope this does not come across in a condescending manner.
Please see the series of posts I made today on the twimlai slack group, on channel #fast_ai_dl, attempting to document items suggested by Jeremy for followup.
We use the gradient of the loss instead of the metric for updating the weights because the loss is better behaved.
More precisely, we use a loss function because it’s differentiable. By using the chain rule, we can propagate its gradient through all the network and update the weights.
Looking for or designing new loss functions for different problems is a research field.
Standard deviation is shift invariant, i.e., std(x) = std(x+c) for any constant c.
See f.e.,
x = np.random.randn(100)
x.std(), (x - 1000).std()
(1.0748678085091383, 1.0748678085091379)
Edit:
Of course if we know what is a distribution of relu’s input, we can rescale it appropriately. If we assume that the input is normally distributed, x ~ normal(0, 1), then:
from sympy import Max, latex # conda install sympy
from sympy.stats import Normal, E, std
X = Normal(‘X’, 0, 1)
reluX = Max(X, 0)
bias = E(reluX)
scale = std(reluX)
# latex(bias), latex(scale)
# float(bias), float(scale)
we get:
\text{bias}=\frac{\sqrt{2}}{2 \sqrt{\pi}} \approx 0.3989\\
\text{scale}\sqrt{- \frac{1}{2 \pi} + \frac{1}{2}} \approx 0.5838
Therefore, with
def renormed_relu(x): return (x.clamp_min(0.) - 0.3989)/0.5838
we should get the output with mean ~= 0.0 and std ~= 1.0. That’s in theory. In practice, lin(x_valid, w1, b1) is not normal(0,1) (more like normal(0.0, 1.4) in my experiments). In this case scale ~= 0.8173 and bias ~= 0.5585 which is much closer to Jeremy’s 0.5 ![]()
I must have pulled in tensor from fastai during my testing at some point. Thanks for the explanation, that was very helpful.
Jeremy said this affected the variance <edit: or std deviation which is its square root>. How can shifting the distribution’s mean affect the variance?
Was that wrong or am I highly confused?
variance is around the mean so it stays the same no matter the offset
Yeah, Jeremy uses it when initializing his weight matrix with this code:
weights = torch.randn(784,10)/math.sqrt(784)
Here is what it does:
Docstring:
randn(*sizes, out=None, dtype=None, layout=torch.strided, device=None, requires_grad=False) -> Tensor
Returns a tensor filled with random numbers from a normal distribution
with mean `0` and variance `1` (also called the standard normal
distribution).
.. math::
\text{out}_{i} \sim \mathcal{N}(0, 1)
The shape of the tensor is defined by the variable argument :attr:`sizes`.
Args:
sizes (int...): a sequence of integers defining the shape of the output tensor.
Can be a variable number of arguments or a collection like a list or tuple.
out (Tensor, optional): the output tensor
dtype (:class:`torch.dtype`, optional): the desired data type of returned tensor.
Default: if ``None``, uses a global default (see :func:`torch.set_default_tensor_type`).
layout (:class:`torch.layout`, optional): the desired layout of returned Tensor.
Default: ``torch.strided``.
device (:class:`torch.device`, optional): the desired device of returned tensor.
Default: if ``None``, uses the current device for the default tensor type
(see :func:`torch.set_default_tensor_type`). :attr:`device` will be the CPU
for CPU tensor types and the current CUDA device for CUDA tensor types.
requires_grad (bool, optional): If autograd should record operations on the
returned tensor. Default: ``False``.
Example::
>>> torch.randn(4)
tensor([-2.1436, 0.9966, 2.3426, -0.6366])
>>> torch.randn(2, 3)
tensor([[ 1.5954, 2.8929, -1.0923],
[ 1.1719, -0.4709, -0.1996]])
Type: builtin_function_or_method
Hi,
Anyone knows where I can find the benchmark code from this High Performance Numeric Programming with Swift: Explorations and Reflections` article?
