Questions or comments about lesson 5? Here’s the place!
The wiki page, including video link, is here: http://wiki.fast.ai/index.php/Lesson_5
Questions or comments about lesson 5? Here’s the place!
The wiki page, including video link, is here: http://wiki.fast.ai/index.php/Lesson_5
I was trying to figure out how the sentiment analysis worked by using much more limited vocabulary from my eldest favorite book green eggs and ham. I did some preprocess in a notebook here and then built a spreadsheet here that made me feel a bit more comfortable that we were convolving words. The activation function at the end isn’t great, but it does give weights that fit a model
I ran into problems then trying to build the model in Keras since I know that my input shape is in the wrong format.
It had been fun to play with.
Very nice! I’m planning to show a similar type of spreadsheet next week. Perhaps I should pick the same corpus as you 
I’m trying to convince myself that that collaborative filtering works as well as I hoped. I’m impressed by the neural net beating the state of the art, but how good is that?
I added a column to the validation set like this:
review = pd.DataFrame(val)
review['prediction'] = nn.predict([val['userId'], val['movieId']])
Eyeballing the result:
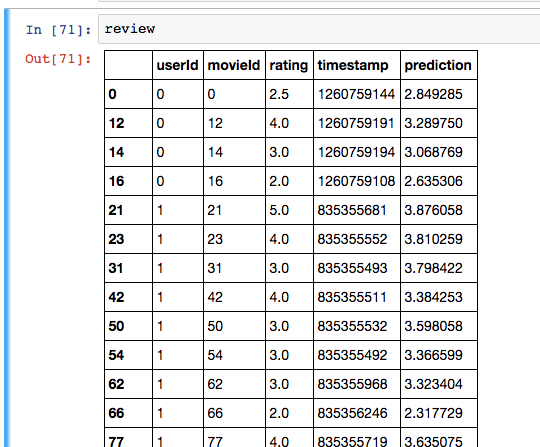
I’m sure there’s a better way to do this:
errors = review.apply(lambda x: np.sqrt(np.abs(np.square(x['rating']) - np.square(x['prediction']))), axis=1)
(np.mean(errors), np.std(errors))
(1.9768158633319872, 0.8483355421950529)
plt.plot(errors)
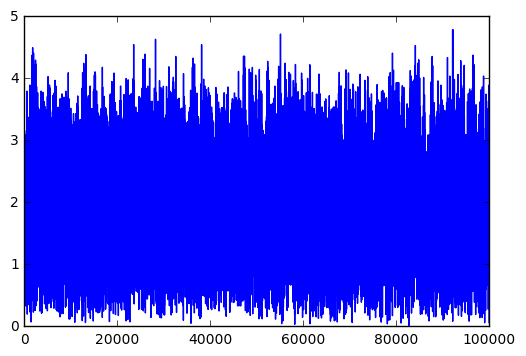
Can you think of a better way to visualize the performance of the model?
Interesting question. The loss function, being mean squared error, is pretty easy to interpret directly. It tells you, on average, how far off the correct rating it was.
You could also round off the ratings and predictions on the validation set to the nearest 0.5, and create a confusion matrix (as we showed in a previous lesson).
Pretty impressive! I was wondering if we can turn this into a classification problem: anyone who has rated 4 or 5 - ‘like/love’ the movie and rating 1 or 2 ‘dislike/hate’ the movie and with a rating of 3 as ‘neutral’. This will offset any chance the model is mostly predicting a rating closer to the mean/median and thereby hiding any flaws. Just a thought.
I was hoping to be able to predict one biomarker, e.g. cortisol, using others, e.g. sodium and potassium. This would be a huge win because sodium and potassium are cheap whereas cortisol is expensive to test. Perhaps collaborative filtering is the wrong tool for the job and I need to sit tight and wait to see what else I discover.
Or maybe it is a classification problem! Maybe I don’t care about the exact number but rather whether cortisol is high or low, and how it compares to baseline over time.
Interesting use case. Maybe have both: classification and regression - provide better guidance or check if they are not contradicting each other.
Do you have a little bit of sample data we could look at?
Of course! Thank you for looking.
The unnormalised data looks like this:
The normalised (sklearn.preprocessing.scale) like this:
As part of the theme of movies and having “silicon valley” consultants in our class: another corpus could be here
I’d guess that a random forest could be a good approach to this - crosstab the table to make each marker a separate column, and the set cortisol as your target variable.
Best I go learn about random forests, thank you!
The last half of this talk will teach you all you need to know! https://www.youtube.com/watch?v=kwt6XEh7U3g
In the single hidden layer NN, the output layer has a size of 500*32 or 16000 neurons. But Keras shows that the number of params is 160000. Where does this 160000 come from?
Here is the picture of the code for quick reference http://wiki.fast.ai/index.php/File:Single_hidden_nn.png
In Lesson 5, imdb dataset classification task (with pretrained vectors, non trainable). I am trying to reproduce the results of the class, but to no avail. The best validation accuracy I can get to is 86%. What is it that I am doing wrong? The model, learning rate etc is exactly the same as that in class.
I’m looking at wordvectors.ipynb
There’s a link “pre-processed glove data”. When I click on the link
I get a 404 Not Found.
Is there an alternate site this has been moved to?
Thanks. --es
BTW. Please put your health before all other things. If you don’t get to this, I’ll try to find another way.
http://www.platform.ai/models/glove/ -> This link worked fine for me.
Thanks.