Thank you for looking and explaining the other pieces.
Thanks @johannesstutz. I’ll take a look at RNNs and see if that clears it up. Appreciate the link to SO post.
Hi all, I’m a little confused with some of content in the MNIST Loss Function section of the lesson.
We first use the linear1 function to apply randomly initialized weights on our training data and then use: ((preds>0.0).float() == train_y).float().mean().item() to calculate an accuracy score for how well the model performed which calculates a score of 0.4912…
I understand this part is more for illustration and is just building up the concepts so they are easy to digest, but am I correct in my perception that this accuracy score isn’t actually measuring anything meaningful? It seems to me that all we have done is applied randomly generated weights to each image (which is just a row with 784 floats representing pixels) and then asked if the scalar output of this is greater than or less than 0. I don’t see why that math would tell us whether the image was a 3 or a 7 and hence comparing that to the labels won’t give us a meaningful answer.
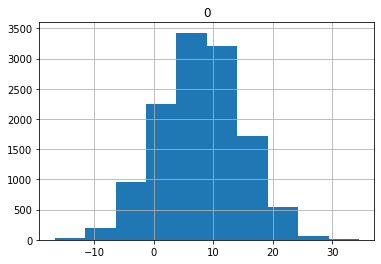
The histogram below shows the distribution of values for my preds tensor. It looks like about 2/3 of the values fall above 0 even though we have a little over 6,000 images of both 3’s and 7’s.
Just to clarify, I understand how the next section works using:
def mnist_loss(predictions, targets): return torch.where(targets==1, 1-predictions, predictions).mean()
This function actually calculates a form of score that is in relation to the target labels and is hence not completely random. I can see how using this along with an optimization step would allow SGD to work.
I just don’t understand how the validation metric (accuracy) is working.
Appreciate any insights on this!
Hi Todd! I’ll try to explain it, I stumbled over this problem as well.
I think the relationship between the activations (predictions) and the labels is just by definition. We have a binary classification task of deciding whether an item is a three or not. So we can just define a prediction of >0 to be a prediction of a three, a prediction of <0 is “not a three” or a seven.
As you correctly mentioned, the weights of the linear layer are randomly generated, which means they don’t have any predictive power before training. In your example, roughly 2/3 of the predictions are above 0 and therefore predict a three. Even if (through a strange weight generation) ALL your predictions where above zero, the accuracy would be around 50% as roughly half of the dataset consists of threes.
After training, the predictions will have a very different distribution. Jumping ahead in the notebook (“Putting it all together”), there is a complete training loop that I used for the following visualizations.
These are the predictions before training, accuracy 52%:
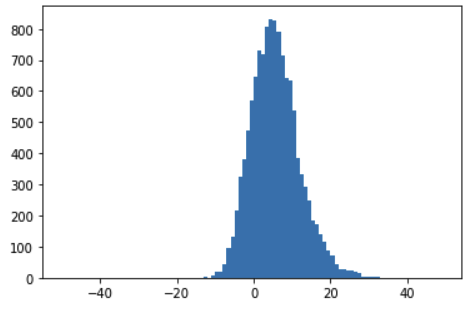
And these are the predictions after training, with an accuracy of 97%:
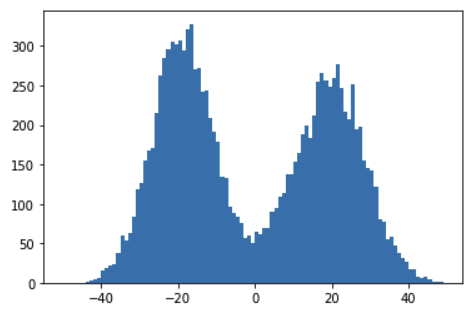
(Notice that I used the raw predictions, without the sigmoid activation function, for better visualization)
I hoped that helped a little!
Thanks for the explanation and second graph, it is now clear to me!
Hey @johannesstutz,
Thanks again for helping me out earler. 
I’ve commented out bits of the code to make it easier to run the whole notebook. Link to notebook here - Lesson 4 Mnist Full
When I train the model, the accuracy does not seem to improve.
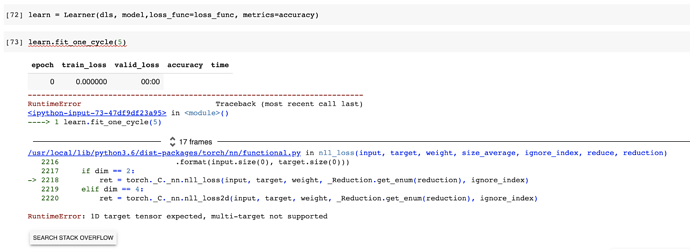
I also tried using the fastai Learner class and ended up with a different error
I’ve tried searching stack overflow and pytorch forums for the source of error and the answers say that the targets need to be a 1D for CrossEntropyLoss. I kind of get that. We need 1 label value for the correct label. Link here Pytorch forum
Is there anything you can see that I’m messing up? Any help is much appreciated!
Adi
Hey Adi, you’re welcome  I’ll have a look at the code again after the weekend. In the meantime, the most important thing - in my experience - is checking the tensor sizes of all inputs and outputs, maybe this will help you get further!
I’ll have a look at the code again after the weekend. In the meantime, the most important thing - in my experience - is checking the tensor sizes of all inputs and outputs, maybe this will help you get further!
Hi everybody.
I have a question about the pixel similarity we used on the MNIST dataset.
I wanted to apply this to our bears example.
So I have been trying to get the image tensors from the DataLoaders (After the DataAugmentation and Cropping has been done) in order to get those to apply pixel similarity on that.
I do know and understand that there really is no point in doing it but I just wanted to do it to see what the results are.
So in short: Is there any way to get the image tensors from DataLoaders
Thanks @johannesstutz!
I’ll definitely try it. That’s a great tip to check the shapes as I go.
In the training dataloader and the test dataloader the shape of a single batch looks like below
This sort of makes sense to me. The batch size is 256 and we have 256 rows. In the input tensor there are 784 pixels and in the target tensor there is 1 label.
The shape of model(xb) and yb are as follows
Again this sort of makes sense to me. There are 10 final activations for the batch of 256 and 1 label for the batch.
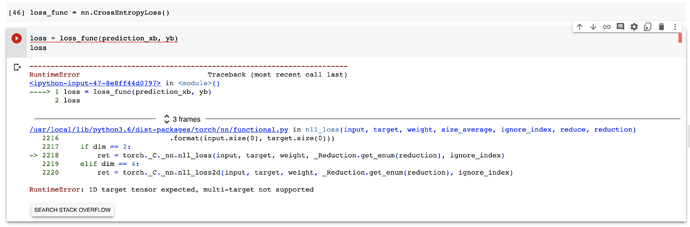
When I tried applying nn.CrossEntropyLoss() on this mini batch just to test it throws an error which suggests the target tensor should be 1D.
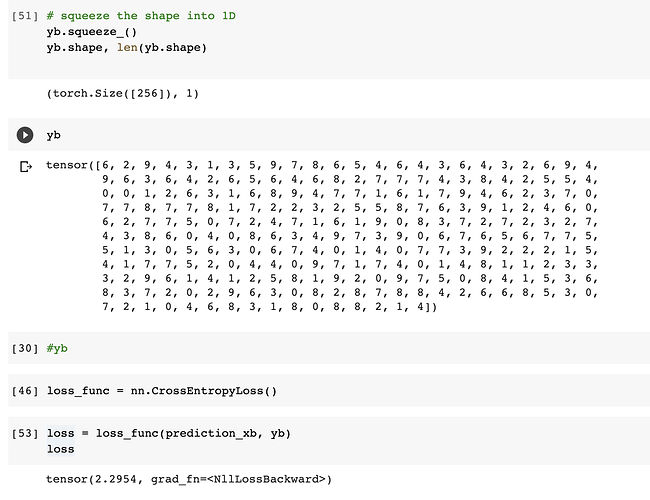
I checked the shape for the yb and it was a 2 dim tensor and the loss func was expecting 1D. I squeezed it in place to convert into 1D. The loss then worked.
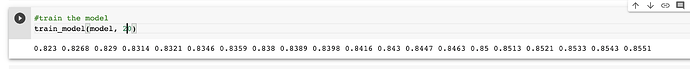
I placed all of this into the usual training loop and it seems to work. It needed about 50 epochs to get to 85% accuracy.
I guess it works?  I am less confused when it comes to the theory but just a bit thrown off that it actually works.
I am less confused when it comes to the theory but just a bit thrown off that it actually works.
I tried using the Learner class as well and it throws the same error around the target needs to be 1D.
I guess I need to try and make the yb of targets into a 1D tensor from a 2D tensor.
Thanks again for your help!
Adi
Congratulations Adi, great work!  I’m glad I could help. Yes, sometimes it feels like magic indeed, when after hours of debugging it “just works”.
I’m glad I could help. Yes, sometimes it feels like magic indeed, when after hours of debugging it “just works”.
For the fastai Learner class, you can try the custom fastai loss function CrossEntropyLossFlat, it flattens the input and target tensors.
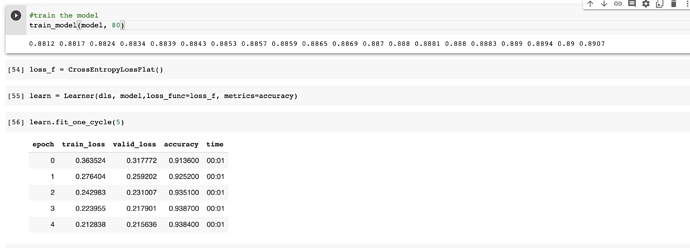
I tried it with CrossEntropyLossFlat and it trained to a higher accuracy in just 5 epochs which is amazing.
Thank you so much! 
I really want to learn this foundational chapter very well before moving ahead because in some ways it feels like most things from here are optimisation tweaks to make training faster and/or with less data.
I am very new to programming and a lot of the programming side of things are confusing or feels like like magic. This forum has been such a help in getting me up to speed.
I’ll be reworking this notebook and try and build as many of these classes and functions from scratch to ensure it makes sense.
Thanks again! 
Perfect 
I think your approach is great, creating all major functions from scratch and making sure to really get the basics right. I should do this more 
And don’t worry about the coding, it will get easier. I had not written a single line of Python before this year and I’m feeling more comfortable every day. I can recommend freecodecamp.org and programiz.com for learning and codewars.com for some fun challenges.
Have fun 
Maybe  You should be able to get the tensors just like this:
You should be able to get the tensors just like this:
xs, ys = first(dls.train)
xs should be a tensor of for example the shape (64, 3, 224, 224) (batch size, channels, image size). Taking the mean over axis 0 gives you the average image in the batch.
You can write a loop over the dataloader like so:
for xs, ys in dls.train:
…
And create the averages for every class.
Good luck 
I solved the MNIST from scratch with the full dataset and my custom learner and loss function. If anyone is interested you can check out the notebook here:
Hi immiemunyi hope your having jolly day.
I had a look at your notebook, great work, nicely written.
Cheers mrfabulous1 

Hello there,
In Chapter 4 in the section An End-to-End SGD Example we define a mean squared error function:
But what I noticed here is that our loss function turns out to be RMSE (because we take a square root in the end).
Is it a mistake in the code? All the following computations look like they were computed using the MSE and not RMSE.
The function
weights*pixelswon’t be flexible enough—it is always equal to 0 when the pixels are equal to 0 (i.e., its intercept is 0).
I don’t really understand - what’s the intuition behind knowing that won’t be flexible enough. Also, why is it equal 0 and why would the pixels be 0?
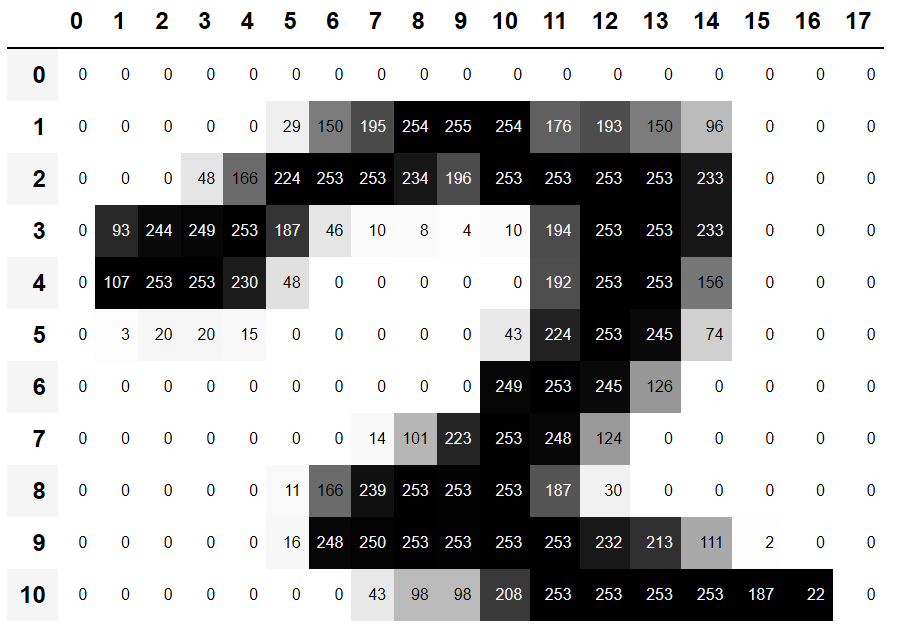
The Pixels would be 0 if the pixel represents a white color (in case of the black and white example we are using.) Remember when we printed out a portion of a sample 3 image and got this:
The pixels that are zero are white and the darkest ones are 255 or closer.
Each pixel will be assigned a weight associated with it. If the input (pixel) is zero, this function weight*pixel will always output a zero, since it is being multiplied with a zero. That’s why its not flexible enough since all the 0 pixels will have zero output. That is why we need a bias, to offset this value since the bias is added. weights*pixels + bias will sort out the issue of a zero pixel.
Thanks for the response! Ok so that makes sense to me, however, I still don’t understand why do we not want 0? seems like that’s still helpful in our loss function? If a pixel is 0 in an eval set but a pixel has a nonnzero value in a test set, can’t we still calculate an error?