Lesson resources
- Video
- Lesson 14 video timeline
- Rossmann data
- Camvid (see Camvid directory)
-
Camvid, additional files (e.g.
label_color.txt)
label_color.txt)I just added videos, notebooks, and data.
Hey @jeremy , the video is only 1hr 15 minutes long. The video runs only up until the break and ends there. 
Wow that’s odd. I have another copy so will upload shortly.
OK I’ve uploaded the full video now. Sorry about that!
the rossman_exp.py file here seems inaccessible…
Me too getting an error with the _exp.py file.
Can’t find rossman_exp.py either. Even looked for rossmann_exp.py (with 2 n’s), couldn’t find that.
Sorry about that! I’ve added that file to part2 github now.
transcript is in the file:
https://drive.google.com/open?id=0BxXRvbqKucuNWEFZbUZwb2RIbFk
in the same directory with all the other transcripts from this course:
https://drive.google.com/open?id=0BxXRvbqKucuNcXdyZ0pYSW5UTzA
please let me know if there are any corrections
and please let me know if there is somewhere else that these could reside …
When I put the MOOC online I’ll make these into youtube captions. 
I am working on the tiramisu-keras notebook, trying to reproduce it.
Sorry if I missed something, but I can’t find the file label_colors.txt
label_codes,label_names = zip(*[
parse_code(l) for l in open(PATH+"label_colors.txt")])
It didn’t come with the above GitHub link to CamVid apparently. I’d be grateful for pointers.
BTW - Nice work on the ```segm_generator````class! Learning a lot from it. And it is useful, too.
Oops sorry! It’s here https://github.com/mostafaizz/camvid
I’m glad! I was pretty pleased with it, I must say ![]()
@jeremy How can we edit/send a push request to the notebooks.
I worked on the rossmann notebook and found a bug:
In Create Models. there are two cells with following content
Cell
map_train = split_cols(cat_map_train) + [contin_map_train]
map_valid = split_cols(cat_map_valid) + [contin_map_valid]
Cell
map_train = split_cols(cat_map_train) + split_cols(contin_map_train)
map_valid = split_cols(cat_map_valid) + split_cols(contin_map_valid)
If I execute it in this order, the 2nd cell will not fit with the format of the model later:
model = Model([inp for inp,emb in embs] + [contin_inp], x)
I need to execute only the first cell and then the format/shape of the arrays fits to the configuration of the model.
The difference is:
When I used the 2nd Cell, the input for the model is too large because the continoues variables are not packed in one array.
I am confused about the labels the model should see at its output.
After a long period of training I got semi-useful predictions, but the model seems to have trouble to get any better and I believe I must habe made a mistake.
BTW - I am training it on a version of PASCAL VOC 2010.
So, the model output looks like
convolution2d_99 (Convolution2D) (None, 256, 256, 22) 5654 merge_101[0][0]
____________________________________________________________________________________________________
reshape_1 (Reshape) (None, 65536, 22) 0 convolution2d_99[0][0]
____________________________________________________________________________________________________
activation_98 (Activation) (None, 65536, 22) 0 reshape_1[0][0]
====================================================================================================
Total params: 9,427,990
Trainable params: 9,324,918
Non-trainable params: 103,072
In my case - 22 classes.
So, the data the segm_generator() spits out is (x, y) with y being the labels of shape (n_samples, rows*cols, 1). So, the classes are encoded into the image as unique integers ranging from 0…21. The model will train.
However, this is nothing like what the Activation layer produces, because it would rather output probabilities for each pixel being of class c between 0 and 1.
So, here my confusion: Even though activation_98 has dims (None, 256256, 22) the model starts fitting with an input of shape (None, 256256, 1) – can this be explained by numpy broadcasting?
However, if I now insert a to_categorical() into segm_generator it’ll output (None, 256*256, 22) one-hot encoded, but the model won’t train. It’ll throw a ValueError:
ValueError: Error when checking model target: expected activation_98 to have shape (None, 65536, 1) but got array with shape (4, 65536, 22)
Why is that?
A related question: In @jeremy’s tiramisu-keras.ipynb notebook there is a whole section on “Convert labels” which I deemed to be necessary for pretty display only, by encoding RGB values in a class dictionary like this.
[((64, 128, 64), 'Animal'),
((192, 0, 128), 'Archway'),
((0, 128, 192), 'Bicyclist'),
((0, 128, 64), 'Bridge'),
((128, 0, 0), 'Building')]
Is that true that it is display only?
Thanks for checking - they’re on github now so can send a PR. Although @bckenstler is heavily changing them for the next couple of weeks.
Although, in general, I don’t expect the notebooks to be things you can just hit shift-enter repeatedly and expect it to work.
That’s because we use sparse_categorical_crossentropy loss, which handles this automatically.
Yeah there’s a couple of different versions of the data I found - not all parts are needed for the dataset I ended up using.
Is anyone else working on the tiramisu semantic segmentation example?
I have been working on training the model on PASCAL VOC 2010 data for a while. I am currently unsure if my model just needs way longer to train or if there is a bug.
Could someone qualitatively judge if the below results make sense?
The input image looks e.g. something like:

The labels look like this (here four labels). Note: This dataset uses one void class which is the outline of the objects.
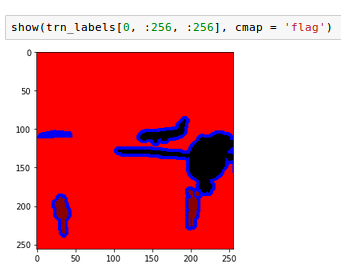
Now, after some 170 epochs (plus minus in different runs) I get a segmentation for the four classes in this picture like:
Top left should be background, top right airplane, bottom left people, bottom right void class (outlines).
Now, while you can say that the model learned something, the result isn’t what it should be. Also, I am starting to wonder if it is a good idea to include the void class at all. Clearly, the model tries to approximate it with a generic edge detector.
The man in the bottom left is almost lost, but he does have poor contrast.
So, essentially I am wondering - is it worth putting more training time in or is it clear (gut-feeling) that there must be a bug?
What makes me a little nervous is that val_acc (and val_loss, not shown) hover around the same value and never seem to improve a lot.
Normalizations I did: Mean subtraction and division by 255.
Labels dataset: At first tried with sparse_cat_crossentropy
However, I started wondering if this is right. With above normalization images scale between [-0.5, 0.5], but the labels are (if one-hot encoded) [0, 1]. At first sight this should still work, but might cause unecessary stress for the model to stretch. So, I am currently trying just division by 255, no mean subtraction.
Really appreciate your opinion.
Some metrics of a typcial training run.
Going back to CamVid this is what I got:
I noticed a few more things:
Rehape, Activation, Reshape or by using hidim_softmax().def hidim_softmax(x, axis=-1):
"""Softmax activation function.
# Arguments
x : Tensor.
axis: Integer, axis along which the softmax normalization is applied.
# Returns
Tensor, output of softmax transformation.
# Raises
ValueError: In case `dim(x) == 1`.
"""
ndim = K.ndim(x)
if ndim == 2:
return K.softmax(x)
elif ndim > 2:
e = K.exp(x - K.max(x, axis=axis, keepdims=True))
s = K.sum(e, axis=axis, keepdims=True)
return e / s
else:
raise ValueError('Cannot apply hidim_softmax to a tensor that is 1D')
The latter is contained in keras 2.0 apparently (I am still on 1.2.2).
Below a modified version of create_tiramisu() found in tiramisu-keras.ipynb. Use either possibility and note you need to modify segm_generator() as well. You need to feed (None, rows, cols, classes) instead of (None, rows*cols, classes).
def create_tiramisu_hd(nb_classes, img_input, nb_dense_block=6,
growth_rate=16, nb_filter=48, nb_layers_per_block=5, p=None, wd=0):
if type(nb_layers_per_block) is list or type(nb_layers_per_block) is tuple:
nb_layers = list(nb_layers_per_block)
else: nb_layers = [nb_layers_per_block] * nb_dense_block
x = conv(img_input, nb_filter, 3, wd, 0)
skips,added = down_path(x, nb_layers, growth_rate, p, wd)
x = up_path(added, reverse(skips[:-1]), reverse(nb_layers[:-1]), growth_rate, p, wd)
x = conv(x, nb_classes, 1, wd, 0)
_,r,c,f = x.get_shape().as_list()
#x = Reshape((-1, nb_classes))(x) # for Activation('softmax')
#x = Activation('softmax')(x) #
#x = Reshape((r, c, f))(x)
x = Lambda(hidim_softmax)(x)
return x