Thanks for your detailed answer Salman,
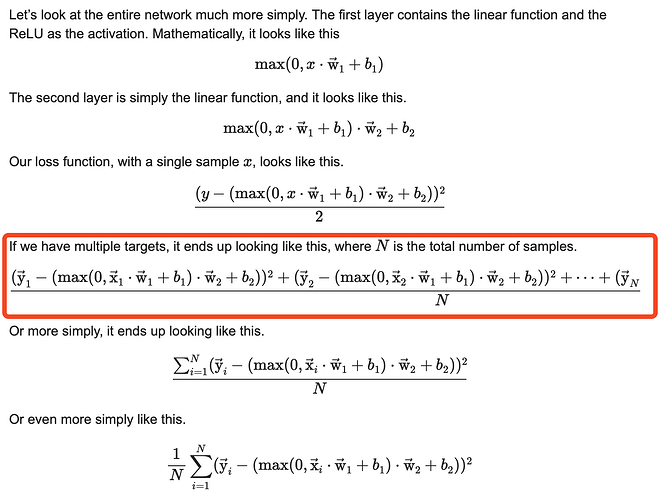
the “hint” guiding the system to the right weights you were writing about in your first answer is what I was referring to as “Yperfect” in my original post - to use the style of your notation I was asking for the origin of yi that you use in your formulas (I think it is called target?) (Btw as you’ve assumed the equation isn’t fully visible, it stops at yn, but the last part with “- max(0,x2⋅w1+b1)⋅w2+b2))^2)” is not visible)
So maybe I can use a little example to describe it in a more precise way what the question from my post was:
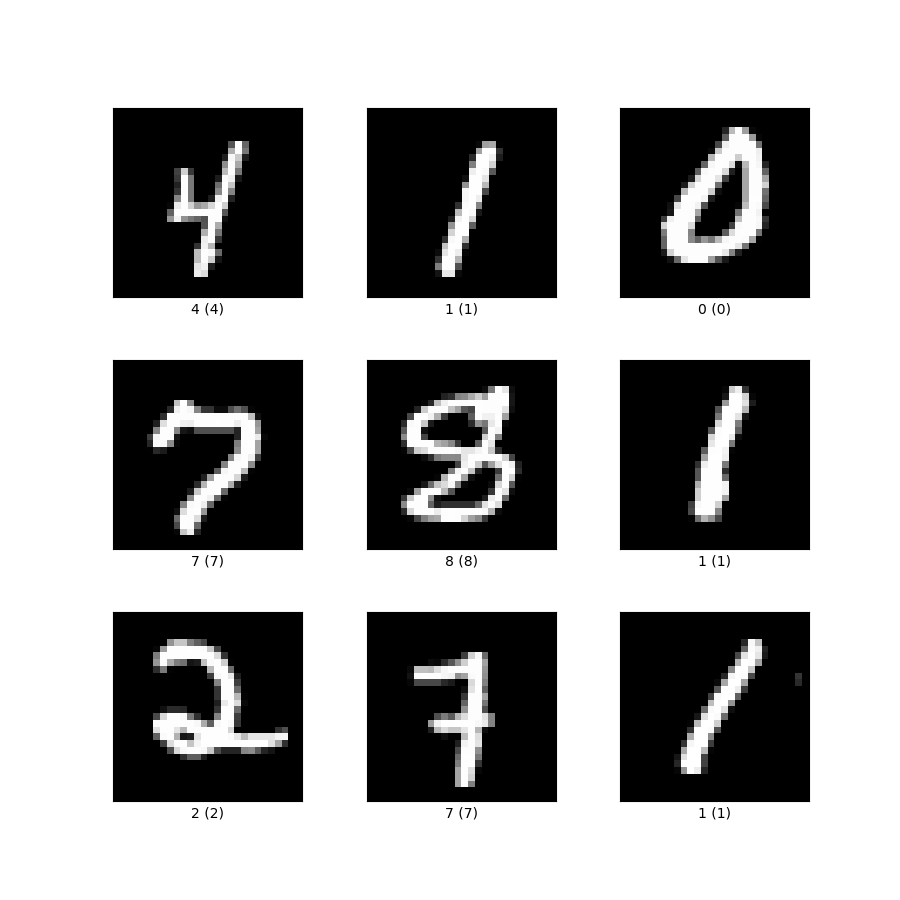
Let’s assume we’d have this input data from the MNIST data set:
So the numbers are:
4, 1, 0, 7, 8, 1, 2, 7, 1
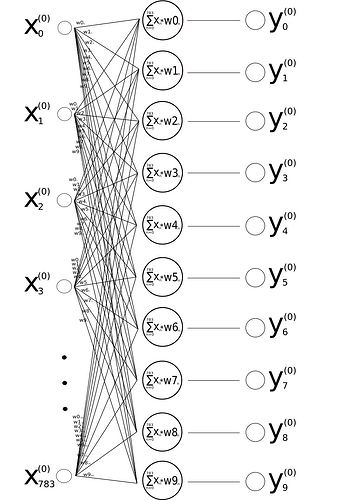
Hence the first line of the matrix X from my original image would be the 28x28 pixel greycale values of the number 4, the second line of number 1, third of number 0 and so on … each of these values multiplied by their weights and summed up would - as drawn in my image - result in the matrix Y (maybe I better should have called it marix A).
Now how do I build the second matrix? Assuming I would change the name of the matrix in my original drawing calling it A instead of Y from now on - how do I get the values for the missing matrix Y?
So would the first three lines of Y then simply be:
0, 0, 0, 0, 1, 0, 0, 0, 0, 0 (number 4)
0, 1, 0, 0, 0, 0, 0, 0, 0, 0 (number 1)
1, 0, 0, 0, 0, 0, 0, 0, 0, 0 (number 0) ?
Is that how you would “encode” the target information into the system so to say?
Using your notation again now we would sum up all the squared subtractions of yi - (max(0,x2⋅w1+b1)⋅w2+b2) - if we would imagine it as an animation line-wise subtracting each vector inside the two matrices line by line - and finally divide the resulting sum by the number of training examples N. Then the systems tries to reduce the error, updates the weights and gets a new error, tries to reduce it, updates the weights and so on.
Or for example in a fashion MNIST case would you then to represent a category also choose one of the 10 columns in the matrix Y? Each category one column in the matrix Y?
Say you’d have to identify 100 different cars, you’d need matrices with 100 columns - is it always like this?
Thanks again for your help, the formulas helped talking about it, very nice of you Salman! Have a good weekend everybody, see forward hearing from you