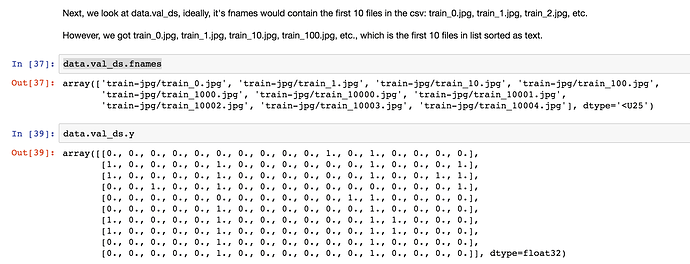
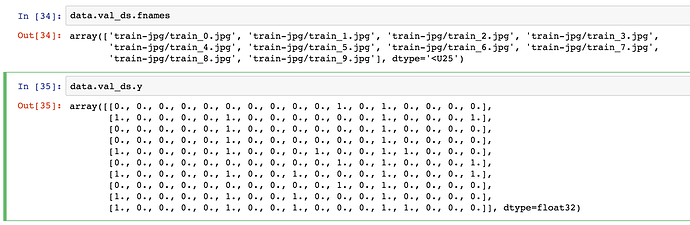
Hi I am using ImageClassifierData.from_csv() to read the data.It has a parameter val_idxs- index of images to be used for validation. e.g. output of get_cv_idxs.If None, default arguments to get_cv_idxs are used.
Problem: I have divided the training dataset in a stratified manner(classification problem) for the purpose of validation data & supplying the original training data indexes of sampled validation data to the val_idxs argument for the data object(ImageClassifierData.from_csv()). I have expected that same validation data as sampled one will be selected via val_idxs.But it returns a different validation data set but with same length(because size is decided by val_idxs size, it is same)
Reasons I could think for the problem:
I think shuffling of the input data is happening first and after it is shuffled the validation indexes provided are selected as validation data in the shuffled data set. Thus a different sample of data is selected from the training data set.
Can I have a way around this. Can I directly give the validation data to the data object.
Thanks in advance for any help/suggestions
Reason found please look into the comments below