Hi everyone,
I am building an application to segment images around people by using COCO dataset and COCO API.
I tried to follow camvid example but you get lost quite soon ![]()
At the moment I can only run the model by keeping one class (which is non sense) and I’m not able to run the model any further to see smart results: probably because I am missing something while building the dataset that gets out later…
I’ll guide you to the problem…
After parsing a while COCO data, I finally had a mask for each file.
By using coco.annToMask I can get mask data and plot it:
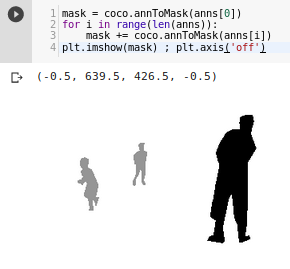
Then I create this function to create images for masks (COCO has masks has annotation in RLE), followed by get_y_fn used later to match each file with its mask:
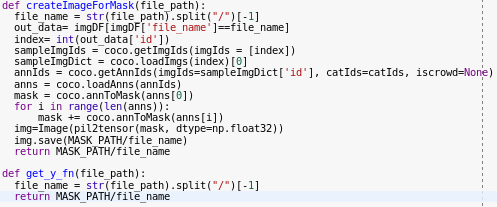
If I use the function I get this:
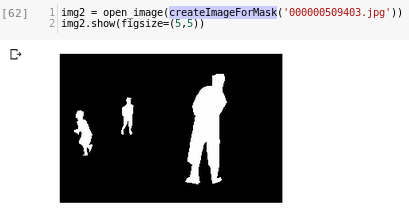
B/W image corresponding to the mask.
First question: is this (an image with 2 colours for a 2-class problem) correct?
I think that original data may be in greyscale because children and adults are mapped in different classes…
Then I try to use fastai open_mask method to see results (this will actually be called from show_batch I guess, in fact those images are blue-ish):
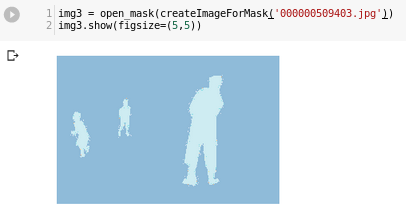
Going on… after building databunch by using COCO API, and running show_batch I can see that data is loaded correctly:

Seems right, right? ![]() So I continued… At first I only kept 1 category “person” and reached the end of learning process, thanks to these lines:
So I continued… At first I only kept 1 category “person” and reached the end of learning process, thanks to these lines:
acc_03 = partial(accuracy_thresh, thresh=0.3)
f_score = partial(fbeta, thresh=0.2, beta = 1)
metrics=[acc_03, f_score]
wd=1e-2
learn = unet_learner(data, models.resnet34, metrics=metrics, wd=wd)
learn.loss_func = MSELossFlat()
Second questions: are these metrics and loss appropriate for this application? Suggestions are appreciated.
By showing results I realized that all images had 1 value predicted (the only class that exists).
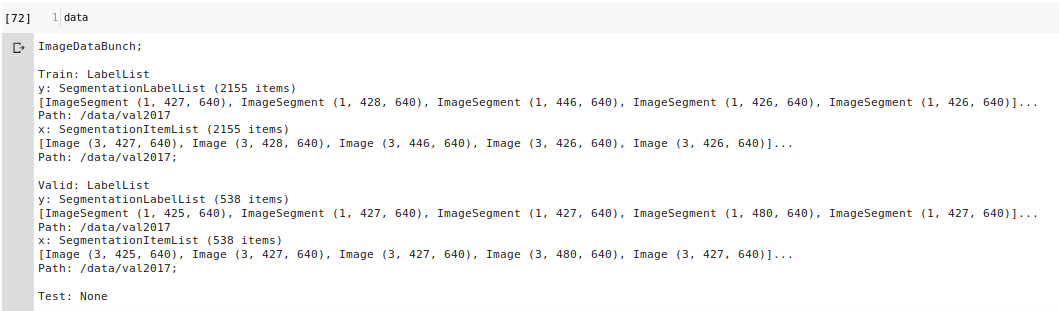
Going back I added to “classes” a new category “background” and checked how classes and dataset were defined:
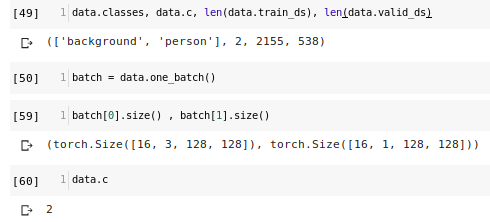
As you can see databunch contains tensors with different number of channels (3 vs 1).
Third question: is this ok? (Since mask is greyscale and original data is RGB. But maybe I missed something)
Other image that shows this difference:
If I try to run the model again I get this error:
The size of tensor a (524288) must match the size of tensor b (262144) at non-singleton dimension 0
On forums and blogs people think is caused by the different number of channels, but I don’t understand why this should be the cause, since the code runs if I keep one class (that has the same number of channels as If I put more classes).
Last question: how are classes and pixel values on mask images joined together?
I think that every colour in grey scale has an index (black: 0, … , white: N) and that is joined to the classes array by index: so first class would be black… is this assumption right? (No clue elsewhere…)
So, what should I do? Change the way I create mask image files? add (how) channels to mask images? have a drink? ![]()
I find it a bit annoying that there is no official tutorial that goes a step beyond the well studied lesson. But this makes things more interesting right?
Anyone happy to help?