Should have thought of that. 
Thanks.
Should have thought of that. 
Thanks.
I think the GeneratorExit error comes from the try-except block in __iter__. Removing the try-except block prevents the GeneratorExit error.
@patch
def __iter__(dl:MixedDL):
"Iterate over your `DataLoader`"
z = zip(*[_loaders[i.fake_l.num_workers==0](i.fake_l) for i in dl.dls])
for b in z:
if dl.device is not None:
b = to_device(b, dl.device)
yield tuple([*dl.dls[0].after_batch(b[0])[:2], *dl.dls[1].after_batch(b[1])])
I think this should work for a test dataset too, I passed an unlabeled dataset to dl.dls[1] and one_batch returned a tuple of length three as expected.
If not, or if you need to do something a little more complicated you can inherit and create a TestMixdDL.
For example I’m currently concatenating multiple labels into one tensor, so the yield line looks like this:
yield tuple([*dl.dls[0].after_batch(b[0])[:2], dl.dls[1].after_batch(b[1][0]),torch.stack(b[1][1:],dim=1)])
and then I have a TestMixedDL that doesn’t return any labels like so:
class TestMixedDL(MixedDL):
def __init__(self, tab_dl:TabDataLoader, vis_dl:TfmdDL, device='cuda:0'):
super().__init__(tab_dl, vis_dl, device)
@patch
def __iter__(dl:TestMixedDL):
"Iterate over your `DataLoader`"
z = zip(*[_loaders[i.fake_l.num_workers==0](i.fake_l) for i in dl.dls])
for b in z:
if dl.device is not None:
b = to_device(b, dl.device)
yield tuple([*dl.dls[0].after_batch(b[0])[:2], dl.dls[1].after_batch(b[1][0])])
I have a separate issue which occurs whether I use the above code or the original with the try-except block. After a period of time, whether it’s a handful of batches or immediately at the start of the second epoch, memory will explode and I will get a CUDA out of memory error. It doesn’t seem to matter how low the batch size is and has happened with a batch size of two.
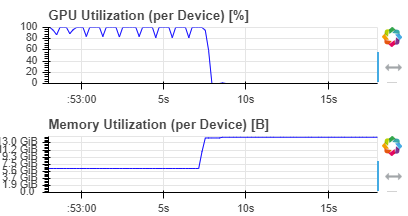
Anyone else running into this issue?
I haven’t quite seen that yet with my text+tab experiments, but what are you using to track the memory usage?
Also how many num_workers are you running?
My images are large: 3D and each is 14mb on disk, which might have something to do with it. I’m using the same number of workers as CPU cores, so that varies on the machine I am using.
The chart is from the Rapids GPU Dashboard:
But the same thing is visible when polling nvidia-smi (both when running in a Jupyter Notebook or JupyterLab), or monitoring the Kaggle GPU usage chart.
It also happens across PyTorch 1.4, 1.5, and 1.5.1.
Hi @muellerzr! Awesome work, I’ll use this on my next project!
I’d like to make a small contribution for learn.show_results() to work:
@patch
def show_results(x:MixedDL,b,out,**kwargs):
for i,dl in enumerate(x.dls):
if i == 0:
dl.show_results(b=b[:2]+(b[3],),out=out,**kwargs)
else:
dl.show_results(b=b[2:],out=out,**kwargs)
@patch
def new(x:MixedDL,*args,**kwargs):
new_dls = [dl.new(*args,**kwargs) for dl in x.dls]
res=MixedDL(*new_dls)
return res
Thank you for sharing this!
Hey this is super helpful!
In terms of training a model with tabular data and images as inputs I guess the two naive routes that I see would be:
-use the embeddings of the vision model as features for the tabular model and train using the tabular model
-use the features from the tabular data as inputs to a vision model that we would most likely concat at the end of the model architecture and train using that modified vision architecture
Am I missing something? Has that been done before? I’ve seen a few threads on how to merge tabular and vision in a smart way, but still not sure of the SOTA way of training such a model.
Thanks!
Elliot
That’s pretty much it. There was a paper published by Facebook about the struggles and how to train such ensemble-based models appropriately, I haven’t looked into it at all yet though.
Thanks for the quick reply and the link to the paper! Pretty interesting paper, might be my next project to try to implement it!
It’s surprising how it seems like such a common problem but without a clear way of solving it (I guess until this paper was published).
Best,
Elliot
This is great Zach!
I got stuck on the same issue as @bwarner, but couldn’t seem to fix it. How would you go about creating mixedDL1 and mixedDL2?
I tried:
mixedDL1 = MixedDL(tab_dl[0], vis_dl[0])
mixedDL2 = MixedDL(tab_dl[1], vis_dl[1])
dls = DataLoaders(mixedDL1, mixedDL2)
learn = cnn_learner(dls, xresnet50, metrics=accuracy)
This returns an error: AttributeError: ‘MixedDL’ object has no attribute ‘after_batch’
You cannot use cnn_learner with this DataLoader as those models won’t work. You need to generate custom architecture. So it’s incomparable there is not fix.
Thank you!
That makes sense.
I would love to see an example of a trained model for this type of problem at some point 
It is in the works. I don’t like non-training examples. But it will be a while
I have a generic version incoming that will be put in as a PR soon. All that’s needed is feeding in your DataLoaders and it will grab the appropriate x’s and y’s (without repeats).
Here’s the big behemoth and all of it’s glory:
class MixedDL():
def __init__(self, *dls, device='cuda:0'):
"Accepts any number of `DataLoaders` and a device"
self.device = device
for dl in dls: dl.shuffle_fn = self.shuffle_fn
self.dls = dls
self.count = 0
self.fake_l = _FakeLoader(self, False, 0, 0)
self._get_idxs()
def __len__(self): return len(self.dls[0])
def _get_vals(self, x):
"Checks for duplicates in batches"
idxs, new_x = [], []
for i, o in enumerate(x): x[i] = o.cpu().numpy().flatten()
for idx, o in enumerate(x):
if not arrayisin(o, new_x):
idxs.append(idx)
new_x.append(o)
return idxs
def _get_idxs(self):
"Get `x` and `y` indicies for batches of data"
dl_dict = dict(zip(range(0,len(self.dls)), [dl.n_inp for dl in self.dls]))
inps = L([])
outs = L([])
for key, n_inp in dl_dict.items():
b = next(iter(self.dls[key]))
inps += L(b[:n_inp])
outs += L(b[n_inp:])
self.x_idxs = self._get_vals(inps)
self.y_idxs = self._get_vals(outs)
def __iter__(self):
z = zip(*[_loaders[i.fake_l.num_workers==0](i.fake_l) for i in self.dls])
for b in z:
inps = []
outs = []
if self.device is not None:
b = to_device(b, self.device)
for batch, dl in zip(b, self.dls):
batch = dl.after_batch(batch)
inps += batch[:dl.n_inp]
outs += batch[dl.n_inp:]
inps = L(inps)[self.x_idxs]
outs = L(outs)[self.y_idxs]
yield (inps, outs)
def one_batch(self):
"Grab one batch of data"
with self.fake_l.no_multiproc(): res = first(self)
if hasattr(self, 'it'): delattr(self, 'it')
return res
def shuffle_fn(self, idxs):
"Shuffle the internal `DataLoaders`"
if self.count == 0:
self.rng = self.dls[0].rng.sample(idxs, len(idxs))
self.count += 1
return self.rng
if self.count == 1:
self.count = 0
return self.rng
def show_batch(self):
"Show a batch of data"
for dl in self.dls:
dl.show_batch()
def to(self, device): self.device = device
And it’s helper:
def _arrayisin(arr, arr_list):
"Checks if `arr` is in `arr_list`"
for a in arr_list:
if np.array_equal(arr, a):
return True
return False
I had to figure out a way to first check how many outputs we had (normally), and then check that all of our y’s are unique, in case we merged two DataLoaders together who both had similar get_y's (or repeated x’s). This is done in the _get_idxs and _get_vals functions.
Awesome job @muellerzr. I have a really dumb question. Can this combination of text and images be done on the Datasets level instead of the Dataloaders level just as you have shown. My idea is to make everything into the datasets class and then use the generic fastai dataloader. It’s just a thought i had
No. We do it at the DataLoader level to avoid headaches of dealing with transforms. It’s a DataLoader of DataLoaders, never interrupting each’s pipeline. (You’d need to do this at that level because of the augmentation pipelines, even tabular has GPU transforms which get the batches)
I understand what you mean. The reason i had this thought is because if i were to do this in normal pytorch code, I’d do it on the dataset level. I still feel the transforms could be applied with typedispatch to the different tensortypes defined in a mixedDatasets and still be able to account for GPU transforms. I also agree that it’ll be a bit of a headache to implement. I’ll still look in to it and update if i have any good results.
I think this approach is a bit complicated defining your own functions. Wouldn’t be better use the DataBlock API ? See for example this solution: https://www.kaggle.com/mnpinto/bengali-ai-fastai2-starter-lb0-9598
I really like the concise code 
I think for this problem (ISIC competition) the code should be something like this:
dblock = DataBlock(
blocks=(ImageBlock(cls=PILDicom), # image_name
CategoryBlock, # sex
CategoryBlock, # anatom_site_general_challenge
RegressionBlock, # age_approx
CategoryBlock)), # target
getters=[ColReader('image_name', pref=train_path, suff='.dcm'),
ColReader('sex'),
ColReader('anatom_site_general_challenge'),
ColReader('age_approx'),
ColReader('target')],
n_inp=4, # Set the number of inputs
item_tfms=Resize(128),
splitter=...,
batch_tfms=...
)
Or use get_x + get_y instead of getters + n_inp.
What do you think about it? How should be the transformation augmentation when you have multiple data types? I’m new to the library, I hope you find this approach interesting.
No. Bengali is fine, they’re all image inputs, the DataBlock API is expected to work as such here. I even made a tutorial notebook myself explaining this. In a multi-modal scenario (what this is designed for), we have multiple different input types. Such as tabular + text, images + tabular + text, so on and so forth. There is not an easy way to bring this into the library, as it involves headaches with the transform pipelines, how do you deal with when you just want to augment your images? How do you make sure your batches all come from the same place? This is what the MixedDL attempts to solve.  What you describe is just a simple scenario where it works
What you describe is just a simple scenario where it works  (While yes technically that is multimodal, it’s a multimodal where the inputs are all the exact same, not what this is designed for). Does this help some?
(While yes technically that is multimodal, it’s a multimodal where the inputs are all the exact same, not what this is designed for). Does this help some? 
Thanks. Yes, headaches with the transform pipelines seems to be the main issue with a single DataBlock. Maybe a list of tmfs (each one for a block) could be a solution for future versions of the library. Here is some pseudocode:
DataBlock(
blocks=[input_dataBlock_1,
input_dataBlock_2,
output_dataBlock_1
output_dataBlock_2],
getters=[getter_input_data_1,
getter_input_data_2,
getter_output_data_1,
getter_output_data_2],
item_tfms=[tmfs_for_input_data_1,
tmfs_for_input_data_2,
tmfs_for_ouput_data_1,
tmfs_for_ouput_data_2]
n_inp=2)
See this thread to why that can be problematic. There’s a lot of workarounds needed here, as the text transforms/API is not the same as the vision, and tabular is a ballpark of its own:
(Notice instead of dealing with the DataBlock API we instead deal with TabularPandas, as tabular operates with this). This method avoids that ones headache, and thanks to the generic method, requires almost no overhead from the user.
If you can find a more successful route please let me know but I’ve been trying to solve this problem for a few months now and this is what I’ve discovered is the best solution. (And Sylvain agrees too)