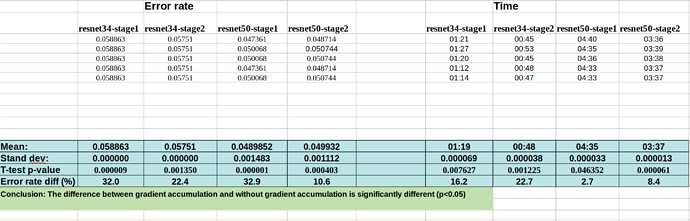
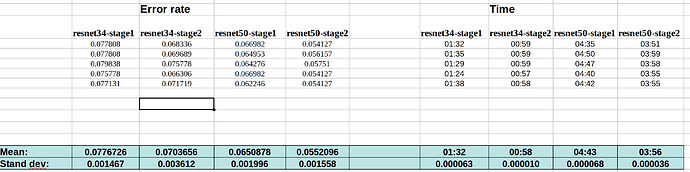
Here is the data for running the FP32 notebooks (with or without gradient accumulation):
Without grad accum.:
With grad accum.:
My intention was to run each notebook 10 times. But from the data, I can see that seed is indeed working perfectly. And there are only tiny variation from run to run within each method.
So the change in the error rate after applying those callbacks is ~25% on average, which cannot be explained by the noise of variation from run to run.
@jeremy
Kerem is going to make a PR request for gradient accumulation with the help of @sgugger . It seems from my analysis above that there is significant difference in the accuracy of doing gradient accumulation of 4 steps with bs=8 (i.e., effectively bs=32), compared to a normal run with bs=32. The difference in error rate is ~25%.
I’ve listed the notebooks used in my previous posts.
Just to be sure before doing the PR, do you think, there is something missing in these callbacks? Isn’t it that we should expect the same error rate in both cases?