Aren’t the models always saved in the same location though? Do you mean it would be nice to be able to choose where they are saved?
Currently they are saved inside tmp\sz\models?
Hi,
In the dataset.py file, we define get_cv_idxs() as:
def get_cv_idxs(n, cv_idx=0, val_pct=0.2, seed=42):
np.random.seed(seed)
n_val = int(val_pct*n)
idx_start = cv_idx*n_val
idxs = np.random.permutation(n)
return idxs[idx_start:idx_start+n_val]
We have set seed=42. Shouldn’t we create truly random idxs everytime we run the code? If the validation loss (or any other metric) is consistent regardless, then, it would mean our model generalizes well.
Many times we care about reproducibility, so it’s useful to have the option. If you want a random value, you could just pass in a value of None for the seed. np.random.seed(None) is Random.
But you bring an important point, may be by default seed should have been None instead of 42. I checked scikit-learn docs and looks like they use None as default for seed. We should also consider changing default value to None.
Yes, this is exactly what I wanted to say  It should be
It should be None by default and the definition should contain the argument seed if the user still wants to set it.
But but 42 gives the answer to everything! 
I would like to add the following to my fastai Wish List:
- print out the names of the metrics
Currently this is the output
A Jupyter Widget
[ 0. 0.03597 0.01879 0.99365]
[ 1. 0.02605 0.01836 0.99365]
[ 2. 0.02189 0.0196 0.99316]
Would be nice if it included the following:
A Jupyter Widget
[epoch training loss validation loss accuracy]
[ 0. 0.03597 0.01879 0.99365]
[ 1. 0.02605 0.01836 0.99365]
[ 2. 0.02189 0.0196 0.99316]
Generally like the approach of having titles to these outputs. The issue is, the last metric could be anything - not always accuracy. So might be better to say it as Val. Metric instead.
@ramesh I just started watching the ML videos, and I’m on Lesson 2, random forests. Do you know what the first two columns represent?
[xxx, xxx, training R^2, validation R^2]
[0.1026724559118164, 0.33553753413792303, 0.9786895444439101, 0.79893791069374753]
Its RMSE for Train and Valiation if this is coming from print_score(m). print_score is defined near the top of the notebook.
Got it! Thank you. @ramesh
For CV I think it’s helpful to have the same split each time.
They are, I believe the point he is making is that, when do you do a relatively large number of cycles with different cycle_len and cycle_mult it is not clear right away after what epochs our model was saved. We do need to make some calculations to find out where our cycles eneded, so I do belive that highlighting in any way the end of the cycle could be very helpful sometimes so that we see what kind of loss and accuracy we get from saved weight right away.
Based on what @KevinB proposed, i suggest something like the following
[ 0. 0.29109 0.22709 0.9116 ] [cycle 0]
[ 1. 0.27596 0.21606 0.91596]
[ 2. 0.25896 0.21201 0.91738] [cycle 1]
[ 3. 0.23578 0.21034 0.91812]
[ 4. 0.25659 0.20687 0.92041]
[ 5. 0.2449 0.1977 0.9226]
[ 6. 0.23827 0.1925 0.92457] [cycle 2]
Yeah thanks for explaining it in a much better way. Usually you can figure out because the loss pops back up but sometimes it’s tough to tell for sure.
Based on what @KevinB proposed, i suggest something like the following
[ 0. 0.29109 0.22709 0.9116 ] [cycle 0]
[ 1. 0.27596 0.21606 0.91596]
[ 2. 0.25896 0.21201 0.91738] [cycle 1]
[ 3. 0.23578 0.21034 0.91812]
[ 4. 0.25659 0.20687 0.92041]
[ 5. 0.2449 0.1977 0.9226]
[ 6. 0.23827 0.1925 0.92457] [cycle 2]
I like that! If anyone wants to try a PR for this I’d be interested - although I’m not quite sure the best design for this. Might be a tricky one!..
Extracting Output from Intermediate Layer (potential feature request)
I am interested in extracting the output of an intermediate layer. For example, the layer right before classification. Is there a relatively straightforward way to do this with Fastai/Pytorch?
I’m thinking I should be able to just refer to the layer name by index and simply call for its output? I’m sure it must be possible but I think it would be nice to have a simple high-level function for doing this. (i.e. model.get_layer(layer_name).output)
Is there a relatively straightforward way to do this with Fastai/Pytorch
Yes, you can use a forward hook. We’ll learn about them on Monday ![]()
Guys,
a very small bug that you must have already seen.
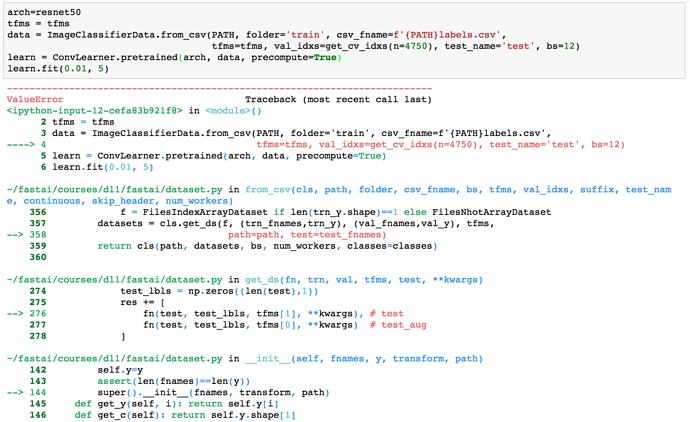
When you define the data object and do provide test_name='test'
data = ImageClassifierData.from_csv(PATH, folder='train', csv_fname=f'{PATH}labels.csv',
tfms=tfms, val_idxs=get_cv_idxs(n=4750), test_name='test', bs=12)
And, there’s no test folder or if the test folder is empty. This assignment results in the following error:
The function read_dir() in dataset.py is used to read the test. The function already contains a TODO: warn or error if no files are found?
def read_dir(path, folder):
# TODO: warn or error if no files found?
full_path = os.path.join(path, folder)
fnames = iglob(f"{full_path}/*.*")
return [os.path.relpath(f,path) for f in fnames]
read_dir() is only used to read the test data folder.
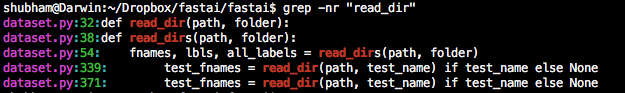
I think this should raise an error is it returns an empty list. Something like the following:
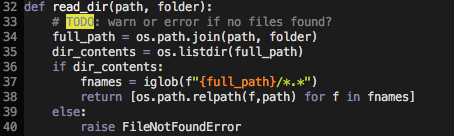
That way the error message would be more explicit.
You can edit your fastai code and check whether the that works and feel free to create a PR, Jeremy will see to it after that…
Yes, I think it’s better to report the issue there and submit a PR.