Wow, the difference is really relevant for Wine and BeetleFly datasets. I assume it is a matter of the super small train size (57 and 20 sequences respectively).
Thanks for your great work @oguiza (enhorabuena!!  )
)
Wow, the difference is really relevant for Wine and BeetleFly datasets. I assume it is a matter of the super small train size (57 and 20 sequences respectively).
Thanks for your great work @oguiza (enhorabuena!!  )
)
Yeah I think working on this subset is a good start.
Good job 
I think it’d also be good to confirm we are not messing up the best perforaming datasets. Here’s the complete list for comparison, including some dataset stats (train size, sequence length, etc).
BTW, the datasets that outperforms HIVE-COTE and TS-CHIEF the most is ChlorineConcentration, which I picked randomly!! 
UCR_SOTAv2.pdf (52.0 KB)
Yeah you are right, we can pick 3 to 4 that would make a fast sanity check.
Just remember that InceptionTime is an ensemble of 5 different Inception architectures, the gain in performance is also attributed to ensembling which reduces the high variance of the individual classifier.
I like the idea of rapid iterations, where we can quickly test many ideas on a small subset of data, so we can discard what doesn’t work quickly. Some of us used this approach with the Imagennete dataset, and the results improved a lot. We can then test the top ones on the entire dataset.
In this case, we could submit the performance for the top 10 underperforming datasets, and the top 3 for example.
I have a list of ideas I’ll test during the week. I’ll post my results. It’ll be important to let the others know what works or doesn’t work, so that we don’t duplicate efforts.
Yeah let’s do that !
@oguiza, i have a bunch of spectra data (XRD) collected in my lab. Simply put each spike in a given data-image corresponds to a mineral and there are libraries for doing this matching… do you think the time series idea can be extended to such dataset. the goal would be to rapid identify the presence/absence of minerals.
Hi @hammao, excellent question!
Although we usually talk about time series, this type of model can actually handle any type of sequential data even if it’s not indexed by time. Actually many of the UCR time series datasets are not time series, because the x axis is not time.
There are examples of spectroscopy (like the OliveOil dataset) where you get a spectrograph of each olive oil sample. In the OliveOil dataset accuracy is around 90%.
I don’t know your dataset or what level of accuracy you need, but I’d definetly give it a try.
You may have seen the timeseriesAI repo I’ve shared. If briefly explain how to prepare the dataset. You can clone it and try it. If you have any questions about how to prepare your data just let me know.
After playing the whole day with LBFGS I am not convinced that it is better than SGD.
For the MLP model, it seems to work sometimes better, but for more complex models it just does not converge to a lower minima. I put a notebook if someone wants to try.
Okay good to know, thanks for sharing.
Thanks for taking the time to test it @tcapelle!
It’s as important to know what works as it is to know what doesn’t work 
I found a difference between our implementations of inception, you have convolutions with bias and I dont’. It appears to work better with bias. I will re post results tomorrow with the corrected Inception implementation, it works way better now, and trains a lot faster. I would say that the bias does not play well with BN but who knows!
I think I got a pretty fastaish implementation of the Inception net:
act_fn = nn.ReLU(inplace=True)
def conv(ni, nf, ks=3, stride=1, bias=False):
return nn.Conv1d(ni, nf, kernel_size=ks, stride=stride, padding=ks//2, bias=bias)
class Shortcut(Module):
"Merge a shortcut with the result of the module by adding them. Adds Conv, BN and ReLU"
def __init__(self, ni, nf, act_fn=act_fn):
self.act_fn=act_fn
self.conv=conv(ni, nf, 1)
self.bn=nn.BatchNorm1d(nf)
def forward(self, x): return act_fn(x + self.bn(self.conv(x.orig)))
class InceptionModule(Module):
"An inception module for TimeSeries, based on https://arxiv.org/pdf/1611.06455.pdf"
def __init__(self, ni, nb_filters=32, kss=[39, 19, 9], bottleneck_size=32, stride=1):
if (bottleneck_size>0 and ni>1): self.bottleneck = conv(ni, bottleneck_size, 1, stride)
else: self.bottleneck = noop
self.convs = nn.ModuleList([conv(bottleneck_size if (bottleneck_size>1 and ni>1) else ni, nb_filters, ks) for ks in listify(kss)])
self.conv_bottle = nn.Sequential(nn.MaxPool1d(3, stride, padding=1), conv(ni, nb_filters, 1))
self.bn_relu = nn.Sequential(nn.BatchNorm1d((len(kss)+1)*nb_filters), nn.ReLU())
def forward(self, x):
return self.bn_relu(torch.cat([c(self.bottleneck(x)) for c in self.convs]+[self.conv_bottle(x)], dim=1))
def create_inception(ni, nout, kss=[39, 19, 9], depth=6, bottleneck_size=32, nb_filters=32, head=True):
"Inception time architecture"
layers = []
n_ks = len(kss) + 1
for d in range(depth):
im = SequentialEx(InceptionModule(1 if d==0 else n_ks*nb_filters, kss=kss, bottleneck_size=bottleneck_size))
if d%3==2: im.append(Shortcut(n_ks*nb_filters, n_ks*nb_filters))
layers.append(im)
head = [AdaptiveConcatPool1d(), Flatten(), nn.Linear(2*4*nb_filters, nout)] if head else []
return nn.Sequential(*layers, *head)
The SequentialEx is a really nice module that let’s you access the input, so just have to append a Shortcut layer et voila, you have a residual block. Another thing I was thinking is that pytorch does not have an equivalent to keras.layers.Concatenate so I made one:
class Cat(Module):
"Concatenate layers outputs over a given dim"
def __init__(self, *layers, dim=1):
self.layers = nn.ModuleList(layers)
self.dim=dim
def forward(self, x):
return torch.cat([l(x) for l in self.layers], dim=self.dim)
note: I am inheriting from fastai Module class, so no __super__ needed.
I’ve now completed 2 tests using fastai.
Both include 3 runs of the bottom10 + top3 as discussed yesterday, with 500 epochs. I run a quick test with 100 and 200 epochs on a couple datasets, and the results improve increaesing the number od eopchs.
It takes about 1.5h to run a single iteration on Google Colab (500 epochs x 13 datasets).
I’ve created a new notebook called 02_UCR_TCS that you can use to run the tests if you want, and make any changes to InceptionTime or the training settings, and it will printout the results for all datasets.
Test 1 result: Baseline
As you’ll see, the end result is a worse than the Fawaz’s InceptionTime single runs, which is a bit dissapointing, but bear in mind there was no hyperparameter tuning.
Test 2 result: Baseline + data augmentation - mixup
Since most datasets are pretty small, I thought it’d be good to apply some data augmentation technique. I have created a version of mixup that we can apply to TS data, which in my opinion works very well. I’m not aware of any paper regarding this, but I think it’s a super powerful technique that usually improves the results reducing overfitting.
When I applied it the results improved, and are very close to the InceptionTime ensemble ones. Bear in mind that there’s no ensemble here, these are just single model results.
I’m preparing a notebook to show you how easily this technique can be used in fastai_timeseries.
I’ve created a measure to be able to compare accuracy of the models, making 100% the best result of HIVE-COTE, TS-CHIEF and InceptionTime.
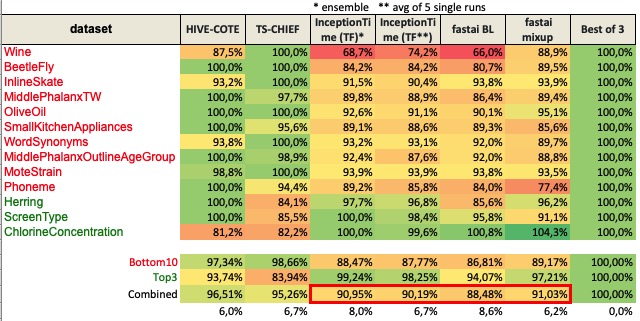
I have a few questions for you @hfawaz:
I think that if we can improve the BL outcome, and have a result closer to the original TensorFlow version, we may have a better result applying mixup1D.
I’ll test other ideas that I have, and will investigate the datasets where mixup didn’t improve data.
Great work.
Yes this is what I am doing for InceptionTime
Yes
I’d need to ckeck it out. I created a cutout and cutmix implementations, and I remember that dor some reason didn’t work out when there was a 3d tensor (time series) instead of a 3d tendor (images) as input. But I can’t remember right now what was the exact change.
@oguiza may be this is a stupid question, but what is the meaning of having an accuracy over 100% in the ChlorineConcentration dataset?
I’ve just briefly reviewed your InceptionTime implementation @tcapelle and the coding style is definitely much better and concise than mine! Great work!
I have a few questions on the implementation and also about the original design.
For @tcapelle:
you don’t seem to use def create_head, you could remove it.
class Shortcut(Module): is duplicated
I hadn’t seen SequentialEx before, but it looks a good way to simplify the code.
I like your Cat class. I’ll use it from now on, referencing you!
I think you are applying different bottleneck layers before each conv_layer. I believe it should only be used once, and then the output goes into all conventional layers.
For @hfawaz:
Do you think that we should benchmark with a maximum number of epochs? I am curious about why you think that one_cycle is not that good. My results with only 40 epochs and one_cycle look reasonably:
| | bl | mixup |
|:-----------------------------|---------:|---------:|
| Wine | 0.5625 | 0.5625 |
| BeetleFly | 0.95 | 0.95 |
| InlineSkate | 0.385455 | 0.354545 |
| MiddlePhalanxTW | 0.616883 | 0.597403 |
| OliveOil | 0.4 | 0.4 |
| SmallKitchenAppliances | 0.789333 | 0.754667 |
| WordSynonyms | 0.65047 | 0.680251 |
| MiddlePhalanxOutlineAgeGroup | 0.655844 | 0.649351 |
| MoteStrain | 0.866613 | 0.890575 |
| Phoneme | 0.227848 | 0.254747 |
| Herring | 0.671875 | 0.625 |
| ScreenType | 0.501333 | 0.501333 |
| ChlorineConcentration | 0.808073 | 0.739844 |
to @oguiza:
mixup is not goodSorry I didn’t explain myself correctly. The 2nd chart with percentages is not accuracy. It’s and index that represents how well a classifier compares to the best of 3 (HIVE-COTE, TS-CHIEF, and InceptionTime). In other words, it’s a way to compare how close it is to the ideas classifier now. Does this help?
That index idea was useful to me to compare performance. Wins/ Losses doesn’t look like a good comparison, and I didn’t feel comfortable with avg accuracy neither.