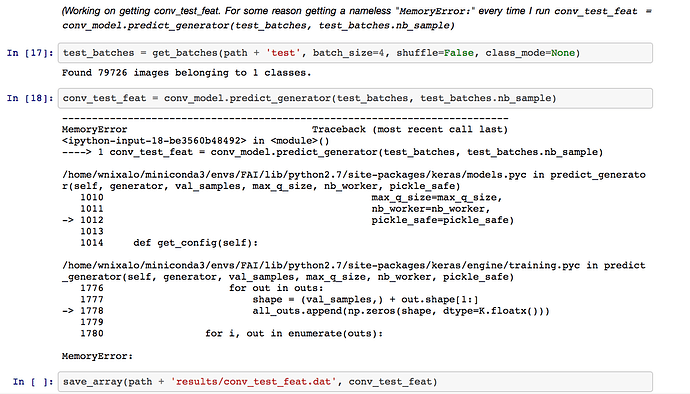
This is great, thanks Bahram. Thing is, this is training the model - I just want to create intermediate features without blowing up the RAM. Looking through documentation & forums now, but haven’t quite found the way how.
So now I know how to create features/predictions on individual or on a specific number of batches:
for batch in test_batches: ...
or
for ...: test_batches.next()
or
xyz = model.predict_generator(test_batches, step)
where step is (I think) a multiple of batch_size, and/or ≥ batch_size and ≤ dataset size.
Now how are these saved to be used for the next step?
I just found out that bcolz saves things in a directory structure, not a single file - so it has a way of keeping track of what’s where. Again, feels like getting too far into the weeds.
There has to be a way to take what @bahram1 said and save features to disk as they’re created.
Update:
Oh hello…
http://bcolz.readthedocs.io/en/latest/reference.html#bcolz.carray.append
append(self, array) Append a numpy array to this instance.
…
http://bcolz.readthedocs.io/en/latest/reference.html#the-carray-class
The carray class
class bcolz.carray
A compressed and enlargeable data container either in-memory or on-disk.
carray exposes a series of methods for dealing with the compressed container in a NumPy-like way.
That sounds a lot like what I was asking for. Will update based on what I find. If anyone has wisdom, please feel free to share.
Update: feels like I’m getting closer.
Same link as above, bcolz.carray class:
rootdir : str, optional
The directory where all the data and metadata will be stored. If specified, then the carray object will be disk-based (i.e. all chunks will live on-disk, not in memory) and persistent (i.e. it can be restored in other session, e.g. via the open() top-level function).
That looks promising. Taking a look at load/save_array in utils.py & the documentation at Library Reference — bcolz 1.2.0 documentation, bcolz.carray.flush() is how bcolz actually saves data to disk. I at first thought it was cleaning a buffer/stream like in C++. Nope.
Furthermore, in the first line of utils.save_array:
bcolz.carray(arr, rootdir=fname, mode='w')
‘w’ erases & overwrites whatever was at fname, but ‘a’ just appends. However that’s for a ‘persistent carray’. Specifying rootdir makes the carray disk-based…
Both are specified in the utils.py implement., so I’m going to guess and say ‘persistent’ isn’t necessarily limited to memory: it just exists… which makes me think… if Howard is using 'w', bcolz isn’t keeping the carray in memory, that’s just convolutional-features variable living in memory. Hopefully carray.flush() doesn’t torpedo this line of thinking, and bcolz is doing some other buffer-witchcraft that doesn’t require keeping everything in memory at once. Fingers crossed.
So, the point?:
Maybe I can use ‘rootdir=..’ and ‘a’ to write my test-convolutional-features to disk using the bcolz carray, as they are created batch by batch. We’ll see.
Update June 9: Done.
Finally got it working; submitted predictions - which also blew my previous best out of the water.
The code to save convolutional features to disk as they are created in batches:
fname = path + 'results/conv_test_feat.dat'
# %rm -r $fname # if you had a previous file there you want to get rid of. (mode='w' would handle that maybe?)
for i in xrange(test_batches.n // beatch_size + 1):
conv_test_feat = conv_model.predict_on_batch(test_batches.next()[0])
if not i:
c = bcolz.carray(conv_feat, rootdir=fname, mode='a')
else:
c.append(conv_test_feat)
c.flush()
The code for generating predictions on the saved convolutions:
idx, inc = 4096, 4096
conv_test_feat = bcolz.open(fname)[:idx]
preds = bn_model.predict(conv_test_feat, batch_size=batch_size, verbose=0)
while idx < test_batches.n - inc:
conv_test_feat = bcolz.open(fname)[idx:idx+inc]
idx += inc
next_preds = bn_model.predict(conv_test-Feat, batch_size=batch_size, verbose=0)
preds = np.concatenate([preds, next_preds])
conv_test_feat = bcolz.open(fname)[idx:]
next_preds = bn_model.predict(conv_test_feat, batch_size=batch_size, verbose=0)
preds = np.concatenate([preds, next_preds])
And that’ll do it. Few notes: of course, this assumes the usual imports, that stuff like test_batches are already defined, and etc. My inability to open an already existing bcolz carray by defining c = bcolz.carray(..), regardless of mode, & success in using c.append(..) after the carray is opened, makes it clear that the first code block can be cleaned up, especially to remove the if-else block. Also, idx and inc, index & increment, are user-defined; I picked them because they seemed big enough to not take too many disk-accesses, but small enough to not put too much into memory at once. Lastly, I take the zeroth index of test_batches.next() because the generator returns as a tuple
Perhaps a few other notes, but that’s all off the top of my head at the moment. I’d love to see a ‘proper/pro’ way to do this (something straight out of keras would be nice!) from J.Howard or someone, but: it works. Looks like it’ll work for big stuff. It’s unconstrained by memory limits (video-mem not-included), so I’m happy with it.
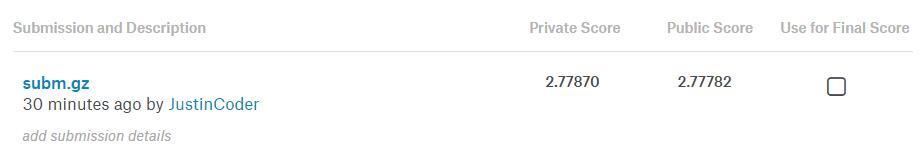
Ah, another note: Doing the above, and running it through the bn_model after training that for I think only 5 epochs (1x 1e-3, 4x 1e-2), got a Kaggle score of 0.70947 at 415/1440 ranking. That’s top 28.9%.
Another thing I haven’t tested is using .fit_generator(..) on conv train/valid features pulled from disk, but that shouldn’t be a huge hassle compared to the above. May update this post down here to include jupyter notebooks for a full implementation, later.
Alright, that’s about it for this one!


 TLDR:
TLDR: