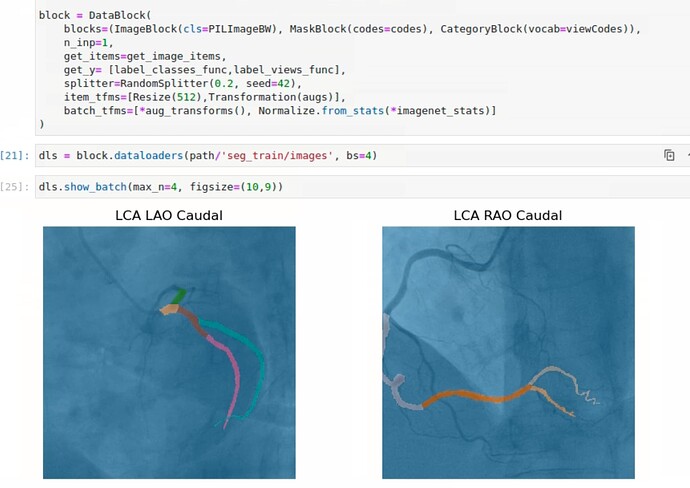
Thank you, @Archaeologist, for your response. Allow me to provide a brief overview of the task at hand. Currently, I am working on predicting coronary artery segments from angiograms using Fastai’s unet_learner, and it has been performing well. While Jeremy’s videos on multi-target learning have been incredibly informative, I encountered a slight difference in my use case, which involves image segmentation rather than classification.
The main idea behind multi-target learning is to improve the model’s predictions across all things it is predicting. Therefore, I decided to explore its potential in our project to enhance segmentation results. To achieve this, I obtained the angles corresponding to the angiogram views and aimed to incorporate them into the unet_learner to predict masks and categories simultaneously. However, it seems that this task is not as straightforward as anticipated, as DynamicUnet doesn’t allow modifications like that. I attempted this by creating a separate class ArteryAndViewClassifier() to configure two heads, but unfortunately, it didn’t work as expected.
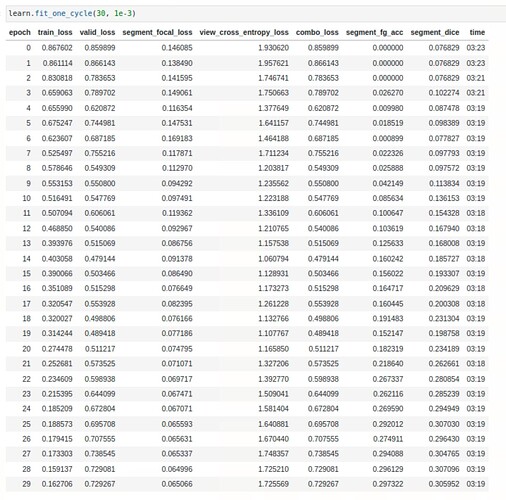
My latest attempt involves utilising the n_out parameter in the unet_learner. By setting it to 37 (26 segmentation classes + 11 view classes), I obtained a bsx37x512x512 output which allowed me to replicate the model’s segmentation part through custom loss and metric functions by utilising the first 26 channels of UNET’s output. However, I am currently facing a challenge with handling the remaining 11 channels in the cross-entropy calculation. My current approach involves summing the entire channel mask to obtain one output per class and then applying sqrt() followed by argmax() to choose one out of the 11 outputs.
Here is the code snippet for the custom loss functions and the combo_loss:
def custom_focal_loss(preds, msk_targs, vu_targs, **kwargs):
return FocalLossFlat(axis=1)(preds[:, :26], msk_targs, **kwargs)
def custom_celf_loss(preds, msk_targs, vu_targs, **kwargs):
cls_targs = targs[:, 26:]
cls_preds = torch.argmax(torch.sqrt(torch.sum(cls_targs, dim=(2, 3))), dim=1)[:, None]
return CrossEntropyLossFlat(reduction='mean')(cls_preds, vu_targs, **kwargs)
def combo_loss(preds, msk_targs, vu_targs, **kwargs):
alpha = 0.5
return (alpha * custom_focal_loss(preds, msk_targs, vu_targs, **kwargs)) + (
(1 - alpha) * custom_celf_loss(preds, msk_targs, vu_targs, **kwargs))
However, I have encountered an error: RuntimeError: “log_softmax_lastdim_kernel_impl” not implemented for ‘Long’. Does this strategy to squeeze the last 26 masks make sense. Any suggestions to resolve this error or implement a better approach.
Thank you again for your valuable help and support!