Hi everyone!
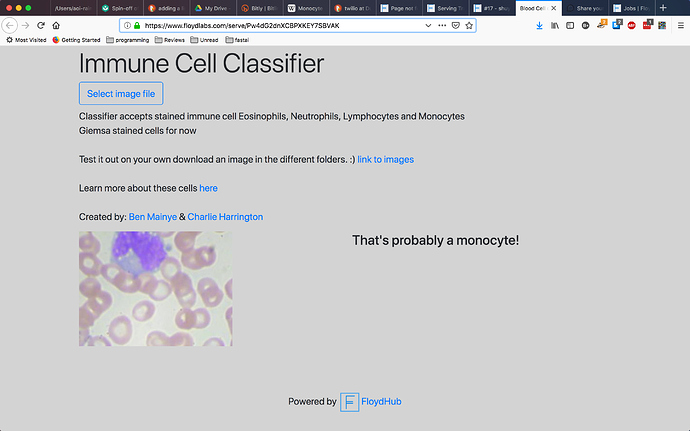
I managed to make an immune cell classifier and I’m serving it at floydhub thanks to @whatrocks. I got my data from Paul Mooney who made it available in kaggle. The model has accuracy of about .95 but I still think I can make it better. Check it out here (Immune cell classifier)[https://www.floydlabs.com/serve/Pw4dG2dnXCBPXKEY7SBVAK] In case I’ve put it offline here’s the screen shot
Sooo excited that the fast.ai library has enabled me to achieve this so quick. It’s been a dream of mine to make this for months. Here’s the repository with code and the model https://github.com/Shuyib/fast-ai-projects