Hi guys!
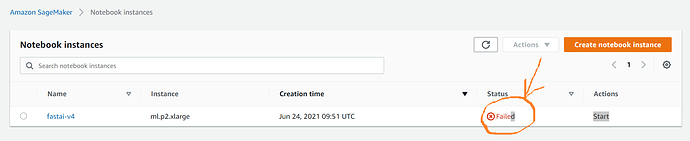
I have faced further issues with running the Notebook instance after applying the initial solution as described above. Specifically, the notebooks are created as expected, however, after stopping the instance (fastai-v4) and coming back to work and restarting it, I would be getting a Failed (to start) error:
Notebook Instance Lifecycle Config ‘arn:aws:sagemaker:us-east-2:255007536151:notebook-instance-lifecycle-config/fastai-v4lifecycleconfig’ for Notebook Instance ‘arn:aws:sagemaker:us-east-2:255007536151:notebook-instance/fastai-v4’ took longer than 5 minutes. Please check your CloudWatch logs for more details if your Notebook Instance has Internet access.
The error in CloudWatch OnStart log exposed that conda install inconsistency persisted if updates are allowed OnStart:
..
==> WARNING: A newer version of conda exists. <==
current version: 4.8.4
latest version: 4.10.1
Please update conda by running
$ conda update -n base -c defaults conda
After further investigation, I wound up editing the FastaiSageMakerStack template (sagemaker-cfn-course-v4.yml) by removing updating conda section in OnStart script section:
templateURL = https://fastai-cfn.s3.amazonaws.com/sagemaker-cfn-course-v4.yml
stackName = FastaiSageMakerStack
This results in updating conda env only after the initial creation (OnCreate) of the Notebook instance (fastai-v4) and not attempting to update it on subsequent starts (OnStart). While this appears to ‘freeze’ the installed conda env version, it has solved the issue.
Here is the code change applied to OnStart section of the template:
original ver (modified per above posts):
echo "Updating conda"
conda update --force-reinstall conda -y
conda update -n base -c defaults conda -y
conda update --all -y
updated version (last three lines removed in OnStart section):
echo "Updating conda - skipped in FastaiSageMakerStack template v6 (sagemaker-cfn-course-v6.yml)"
Below is the updated ver (sagemaker-cfn-course-v4_updated.yml) of the stack template to install the stack from scratch instead of creating the stack from the templates listed here: Amazon SageMaker | Practical Deep Learning for Coders (which results in the error).
To create the stack from scratch, save the template below as sagemaker-cfn-course-v4_updated.yml file. Then delete the failing stack and create it from scratch from this template.
Checking the CloudWatch logs (OnCreate and OnStart) should reflect now errors and successful avoiding of attempting to update conda OnStart
Parameters:
InstanceType:
Type: String
Default: ml.p2.xlarge
AllowedValues:
- ml.p3.2xlarge
- ml.p2.xlarge
Description: Enter the SageMaker Notebook instance type
VolumeSize:
Type: Number
Default: 50
Description: Enter the size of the EBS volume attached to the notebook instance
MaxValue: 17592
MinValue: 5
Resources:
Fastai2SagemakerNotebookfastaiv4NotebookRoleA75B4C74:
Type: AWS::IAM::Role
Properties:
AssumeRolePolicyDocument:
Statement:
- Action: sts:AssumeRole
Effect: Allow
Principal:
Service: sagemaker.amazonaws.com
Version: "2012-10-17"
ManagedPolicyArns:
- Fn::Join:
- ""
- - "arn:"
- Ref: AWS::Partition
- :iam::aws:policy/AmazonSageMakerFullAccess
Metadata:
aws:cdk:path: CdkFastaiv2SagemakerNbStack/Fastai2SagemakerNotebook/fastai-v4NotebookRole/Resource
Fastai2SagemakerNotebookfastaiv4LifecycleConfigD72E2247:
Type: AWS::SageMaker::NotebookInstanceLifecycleConfig
Properties:
NotebookInstanceLifecycleConfigName: fastai-v4LifecycleConfig
OnCreate:
- Content:
Fn::Base64: >-
#!/bin/bash
set -e
echo "Starting on Create script"
sudo -i -u ec2-user bash <<EOF
touch /home/ec2-user/SageMaker/.create-notebook
EOF
cat > /home/ec2-user/SageMaker/.fastai-install.sh <<\EOF
#!/bin/bash
set -e
echo "Creating dirs and symlinks"
mkdir -p /home/ec2-user/SageMaker/.cache
mkdir -p /home/ec2-user/SageMaker/.fastai
[ ! -L "/home/ec2-user/.cache" ] && ln -s /home/ec2-user/SageMaker/.cache /home/ec2-user/.cache
[ ! -L "/home/ec2-user/.fastai" ] && ln -s /home/ec2-user/SageMaker/.fastai /home/ec2-user/.fastai
echo "Updating conda"
conda update --force-reinstall conda -y
conda update -n base -c defaults conda -y
conda update --all -y
echo "Starting conda create command for fastai env"
conda create -mqyp /home/ec2-user/SageMaker/.env/fastai python=3.6
echo "Activate fastai conda env"
conda init bash
source ~/.bashrc
conda activate /home/ec2-user/SageMaker/.env/fastai
echo "Install ipython kernel and widgets"
conda install ipywidgets ipykernel -y
echo "Installing fastai lib"
pip install -r /home/ec2-user/SageMaker/fastbook/requirements.txt
pip install fastbook sagemaker
echo "Installing Jupyter kernel for fastai"
python -m ipykernel install --name 'fastai' --user
echo "Finished installing fastai conda env"
echo "Install Jupyter nbextensions"
conda activate JupyterSystemEnv
pip install jupyter_contrib_nbextensions
jupyter contrib nbextensions install --user
echo "Restarting jupyter notebook server"
pkill -f jupyter-notebook
rm /home/ec2-user/SageMaker/.create-notebook
echo "Exiting install script"
EOF
chown ec2-user:ec2-user /home/ec2-user/SageMaker/.fastai-install.sh
chmod 755 /home/ec2-user/SageMaker/.fastai-install.sh
sudo -i -u ec2-user bash <<EOF
nohup /home/ec2-user/SageMaker/.fastai-install.sh &
EOF
echo "Finishing on Create script"
OnStart:
- Content:
Fn::Base64: >-
#!/bin/bash
set -e
echo "Starting on Start script"
sudo -i -u ec2-user bash << EOF
if [[ -f /home/ec2-user/SageMaker/.create-notebook ]]; then
echo "Skipping as currently installing conda env"
else
# create symlinks to EBS volume
echo "Creating symlinks"
ln -s /home/ec2-user/SageMaker/.fastai /home/ec2-user/.fastai
echo "Updating conda - skipped in FastaiSageMakerStack template v4_updated (sagemaker-cfn-course-v4_updated.yml)"
echo "Activate fastai conda env"
conda init bash
source ~/.bashrc
conda activate /home/ec2-user/SageMaker/.env/fastai
echo "Updating fastai packages"
pip install fastai fastcore sagemaker --upgrade
echo "Installing Jupyter kernel"
python -m ipykernel install --name 'fastai' --user
echo "Install Jupyter nbextensions"
conda activate JupyterSystemEnv
pip install jupyter_contrib_nbextensions
jupyter contrib nbextensions install --user
echo "Restarting jupyter notebook server"
pkill -f jupyter-notebook
echo "Finished setting up Jupyter kernel"
fi
EOF
echo "Finishing on Start script"
Metadata:
aws:cdk:path: CdkFastaiv2SagemakerNbStack/Fastai2SagemakerNotebook/fastai-v4LifecycleConfig
Fastai2SagemakerNotebookfastaiv4NotebookInstance7C46E7E0:
Type: AWS::SageMaker::NotebookInstance
Properties:
InstanceType:
Ref: InstanceType
RoleArn:
Fn::GetAtt:
- Fastai2SagemakerNotebookfastaiv4NotebookRoleA75B4C74
- Arn
DefaultCodeRepository: https://github.com/fastai/fastbook
LifecycleConfigName: fastai-v4LifecycleConfig
NotebookInstanceName: fastai-v4
VolumeSizeInGB:
Ref: VolumeSize
Metadata:
aws:cdk:path: CdkFastaiv2SagemakerNbStack/Fastai2SagemakerNotebook/fastai-v4NotebookInstance
CDKMetadata:
Type: AWS::CDK::Metadata
Properties:
Modules: aws-cdk=1.60.0,@aws-cdk/aws-iam=1.60.0,@aws-cdk/aws-sagemaker=1.60.0,@aws-cdk/cloud-assembly-schema=1.60.0,@aws-cdk/core=1.60.0,@aws-cdk/cx-api=1.60.0,@aws-cdk/region-info=1.60.0,jsii-runtime=node.js/v14.8.0
Condition: CDKMetadataAvailable
Conditions:
CDKMetadataAvailable:
Fn::Or:
- Fn::Or:
- Fn::Equals:
- Ref: AWS::Region
- ap-east-1
- Fn::Equals:
- Ref: AWS::Region
- ap-northeast-1
- Fn::Equals:
- Ref: AWS::Region
- ap-northeast-2
- Fn::Equals:
- Ref: AWS::Region
- ap-south-1
- Fn::Equals:
- Ref: AWS::Region
- ap-southeast-1
- Fn::Equals:
- Ref: AWS::Region
- ap-southeast-2
- Fn::Equals:
- Ref: AWS::Region
- ca-central-1
- Fn::Equals:
- Ref: AWS::Region
- cn-north-1
- Fn::Equals:
- Ref: AWS::Region
- cn-northwest-1
- Fn::Equals:
- Ref: AWS::Region
- eu-central-1
- Fn::Or:
- Fn::Equals:
- Ref: AWS::Region
- eu-north-1
- Fn::Equals:
- Ref: AWS::Region
- eu-west-1
- Fn::Equals:
- Ref: AWS::Region
- eu-west-2
- Fn::Equals:
- Ref: AWS::Region
- eu-west-3
- Fn::Equals:
- Ref: AWS::Region
- me-south-1
- Fn::Equals:
- Ref: AWS::Region
- sa-east-1
- Fn::Equals:
- Ref: AWS::Region
- us-east-1
- Fn::Equals:
- Ref: AWS::Region
- us-east-2
- Fn::Equals:
- Ref: AWS::Region
- us-west-1
- Fn::Equals:
- Ref: AWS::Region
- us-west-2
I hope this helps.
Best,
PO