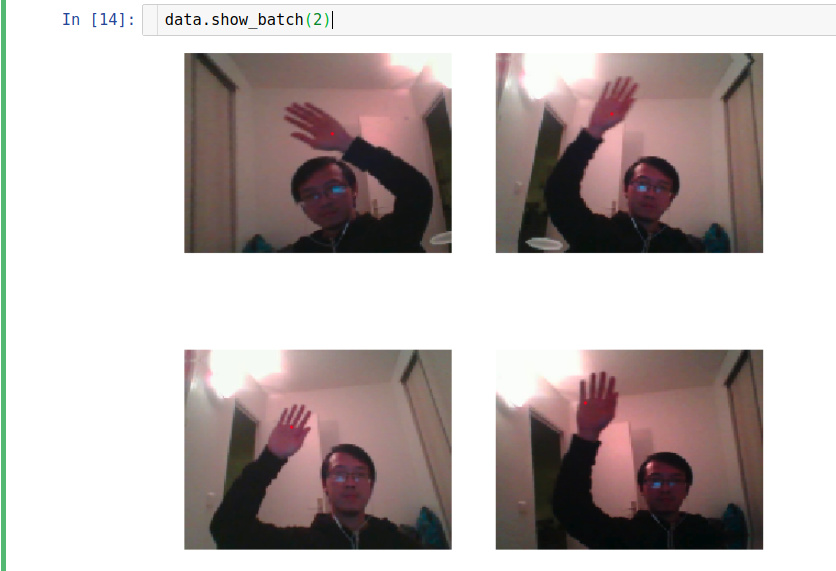
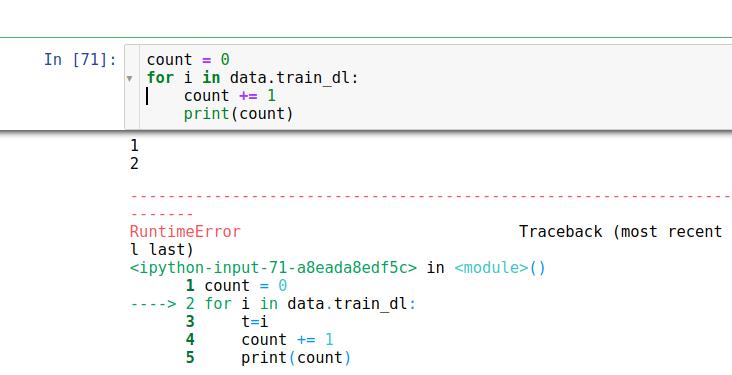
I am trying to create data with PointsItemList for a hand tracking problem. The code seems correct that data.show_batch show exactly what I want.
data = (PointsItemList.from_df(df, path=path, folder='train', cols=['frame'], suffix='.png')
.random_split_by_pct()
.label_from_df(cols=['loc'])
.transform(get_transforms(), tfm_y=True, size=(120,160))
.databunch().normalize()
)
However learn.lr_find() or learn.fit() randomly have the error below
---------------------------------------------------------------------------
RuntimeError Traceback (most recent call last)
<ipython-input-24-4dfb24161c57> in <module>()
----> 1 learn.fit_one_cycle(1)
~/fastai/fastai/train.py in fit_one_cycle(learn, cyc_len, max_lr, moms, div_factor, pct_start, wd, callbacks, **kwargs)
20 callbacks.append(OneCycleScheduler(learn, max_lr, moms=moms, div_factor=div_factor,
21 pct_start=pct_start, **kwargs))
---> 22 learn.fit(cyc_len, max_lr, wd=wd, callbacks=callbacks)
23
24 def lr_find(learn:Learner, start_lr:Floats=1e-7, end_lr:Floats=10, num_it:int=100, stop_div:bool=True, **kwargs:Any):
~/fastai/fastai/basic_train.py in fit(self, epochs, lr, wd, callbacks)
164 callbacks = [cb(self) for cb in self.callback_fns] + listify(callbacks)
165 fit(epochs, self.model, self.loss_func, opt=self.opt, data=self.data, metrics=self.metrics,
--> 166 callbacks=self.callbacks+callbacks)
167
168 def create_opt(self, lr:Floats, wd:Floats=0.)->None:
~/fastai/fastai/basic_train.py in fit(epochs, model, loss_func, opt, data, callbacks, metrics)
92 except Exception as e:
93 exception = e
---> 94 raise e
95 finally: cb_handler.on_train_end(exception)
96
~/fastai/fastai/basic_train.py in fit(epochs, model, loss_func, opt, data, callbacks, metrics)
80 cb_handler.on_epoch_begin()
81
---> 82 for xb,yb in progress_bar(data.train_dl, parent=pbar):
83 xb, yb = cb_handler.on_batch_begin(xb, yb)
84 loss = loss_batch(model, xb, yb, loss_func, opt, cb_handler)
~/anaconda3/lib/python3.6/site-packages/fastprogress/fastprogress.py in __iter__(self)
63 self.update(0)
64 try:
---> 65 for i,o in enumerate(self._gen):
66 yield o
67 if self.auto_update: self.update(i+1)
~/fastai/fastai/basic_data.py in __iter__(self)
68 def __iter__(self):
69 "Process and returns items from `DataLoader`."
---> 70 for b in self.dl:
71 #y = b[1][0] if is_listy(b[1]) else b[1] # XXX: Why is this line here?
72 yield self.proc_batch(b)
~/anaconda3/lib/python3.6/site-packages/torch/utils/data/dataloader.py in __next__(self)
337 if self.rcvd_idx in self.reorder_dict:
338 batch = self.reorder_dict.pop(self.rcvd_idx)
--> 339 return self._process_next_batch(batch)
340
341 if self.batches_outstanding == 0:
~/anaconda3/lib/python3.6/site-packages/torch/utils/data/dataloader.py in _process_next_batch(self, batch)
372 self._put_indices()
373 if isinstance(batch, ExceptionWrapper):
--> 374 raise batch.exc_type(batch.exc_msg)
375 return batch
376
RuntimeError: Traceback (most recent call last):
File "/home/hoatruong/anaconda3/lib/python3.6/site-packages/torch/utils/data/dataloader.py", line 114, in _worker_loop
samples = collate_fn([dataset[i] for i in batch_indices])
File "/home/hoatruong/fastai/fastai/torch_core.py", line 105, in data_collate
return torch.utils.data.dataloader.default_collate(to_data(batch))
File "/home/hoatruong/anaconda3/lib/python3.6/site-packages/torch/utils/data/dataloader.py", line 198, in default_collate
return [default_collate(samples) for samples in transposed]
File "/home/hoatruong/anaconda3/lib/python3.6/site-packages/torch/utils/data/dataloader.py", line 198, in <listcomp>
return [default_collate(samples) for samples in transposed]
File "/home/hoatruong/anaconda3/lib/python3.6/site-packages/torch/utils/data/dataloader.py", line 175, in default_collate
return torch.stack(batch, 0, out=out)
RuntimeError: invalid argument 0: Sizes of tensors must match except in dimension 0. Got 1 and 0 in dimension 1 at /opt/conda/conda-bld/pytorch-nightly_1538905146867/work/aten/src/TH/generic/THTensorMoreMath.cpp:1317
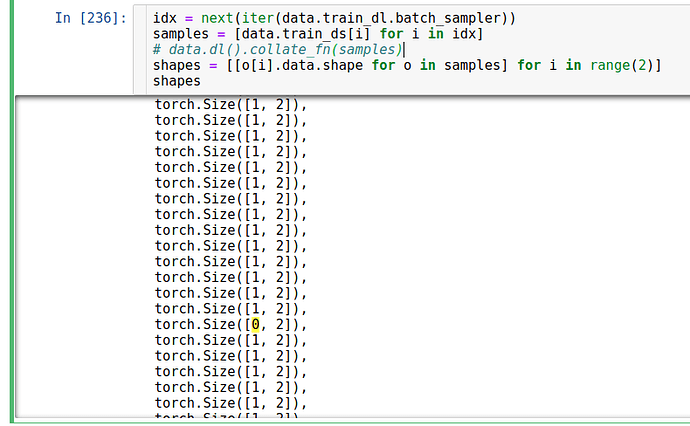
I was thinking that the shape of every elements in data are not coherent but it is not.
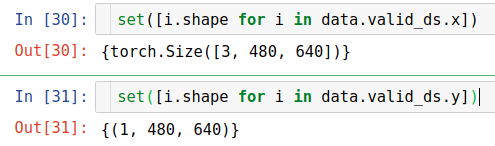
I found that the error might come from the data_loader when we grab a new batch. But it is quite randomly that the order when I get the error is not the same.
I hope someone can help me to clarify this problem. Thank you so much in advance