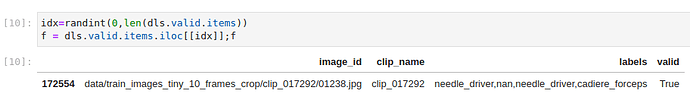
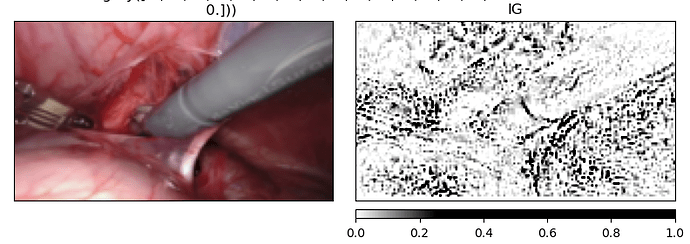
Thanks. it worked. Now the code is picking a row from the data frame.
But now stumbled upon ‘wrong size in the last dimension error’. Please check below:
RuntimeError Traceback (most recent call last)
Cell In [7], line 5
2 f = dls.valid.items.iloc[[idx]]
4 captum=CaptumInterpretation(learn)
----> 5 captum.visualize(f)
File ~/mambaforge/lib/python3.10/site-packages/fastai/callback/captum.py:55, in CaptumInterpretation.visualize(self, inp, metric, n_steps, baseline_type, nt_type, strides, sliding_window_shapes)
53 tls = L([TfmdLists(inp, t) for t in L(ifnone(self.dls.tfms,[None]))])
54 inp_data=list(zip(*(tls[0],tls[1])))[0]
—> 55 enc_data,dec_data=self._get_enc_dec_data(inp_data)
56 attributions=self._get_attributions(enc_data,metric,n_steps,nt_type,baseline_type,strides,sliding_window_shapes)
57 self._viz(attributions,dec_data,metric)
File ~/mambaforge/lib/python3.10/site-packages/fastai/callback/captum.py:73, in CaptumInterpretation._get_enc_dec_data(self, inp_data)
71 def _get_enc_dec_data(self,inp_data):
72 dec_data=self.dls.after_item(inp_data)
—> 73 enc_data=self.dls.after_batch(to_device(self.dls.before_batch(dec_data),self.dls.device))
74 return(enc_data,dec_data)
File ~/mambaforge/lib/python3.10/site-packages/fastcore/transform.py:208, in Pipeline.call(self, o)
→ 208 def call(self, o): return compose_tfms(o, tfms=self.fs, split_idx=self.split_idx)
File ~/mambaforge/lib/python3.10/site-packages/fastcore/transform.py:158, in compose_tfms(x, tfms, is_enc, reverse, **kwargs)
156 for f in tfms:
157 if not is_enc: f = f.decode
→ 158 x = f(x, **kwargs)
159 return x
File ~/mambaforge/lib/python3.10/site-packages/fastai/vision/augment.py:49, in RandTransform.call(self, b, split_idx, **kwargs)
43 def call(self,
44 b,
45 split_idx:int=None, # Index of the train/valid dataset
46 **kwargs
47 ):
48 self.before_call(b, split_idx=split_idx)
—> 49 return super().call(b, split_idx=split_idx, **kwargs) if self.do else b
File ~/mambaforge/lib/python3.10/site-packages/fastcore/transform.py:81, in Transform.call(self, x, **kwargs)
—> 81 def call(self, x, **kwargs): return self._call(‘encodes’, x, **kwargs)
File ~/mambaforge/lib/python3.10/site-packages/fastcore/transform.py:91, in Transform._call(self, fn, x, split_idx, **kwargs)
89 def _call(self, fn, x, split_idx=None, **kwargs):
90 if split_idx!=self.split_idx and self.split_idx is not None: return x
—> 91 return self._do_call(getattr(self, fn), x, **kwargs)
File ~/mambaforge/lib/python3.10/site-packages/fastcore/transform.py:98, in Transform.do_call(self, f, x, **kwargs)
96 ret = f.returns(x) if hasattr(f,‘returns’) else None
97 return retain_type(f(x, **kwargs), x, ret)
—> 98 res = tuple(self.do_call(f, x, **kwargs) for x in x)
99 return retain_type(res, x)
File ~/mambaforge/lib/python3.10/site-packages/fastcore/transform.py:98, in (.0)
96 ret = f.returns(x) if hasattr(f,‘returns’) else None
97 return retain_type(f(x, **kwargs), x, ret)
—> 98 res = tuple(self.do_call(f, x, **kwargs) for x_ in x)
99 return retain_type(res, x)
File ~/mambaforge/lib/python3.10/site-packages/fastcore/transform.py:97, in Transform.do_call(self, f, x, **kwargs)
95 if f is None: return x
96 ret = f.returns(x) if hasattr(f,‘returns’) else None
—> 97 return retain_type(f(x, **kwargs), x, ret)
98 res = tuple(self.do_call(f, x, **kwargs) for x in x)
99 return retain_type(res, x)
File ~/mambaforge/lib/python3.10/site-packages/fastcore/dispatch.py:120, in TypeDispatch.call(self, *args, **kwargs)
118 elif self.inst is not None: f = MethodType(f, self.inst)
119 elif self.owner is not None: f = MethodType(f, self.owner)
→ 120 return f(*args, **kwargs)
File ~/mambaforge/lib/python3.10/site-packages/fastai/vision/augment.py:501, in AffineCoordTfm.encodes(self, x)
→ 501 def encodes(self, x:TensorImage): return self._encode(x, self.mode)
File ~/mambaforge/lib/python3.10/site-packages/fastai/vision/augment.py:499, in AffineCoordTfm._encode(self, x, mode, reverse)
497 def _encode(self, x, mode, reverse=False):
498 coord_func = None if len(self.coord_fs)==0 or self.split_idx else partial(compose_tfms, tfms=self.coord_fs, reverse=reverse)
→ 499 return x.affine_coord(self.mat, coord_func, sz=self.size, mode=mode, pad_mode=self.pad_mode, align_corners=self.align_corners)
File ~/mambaforge/lib/python3.10/site-packages/fastai/vision/augment.py:391, in affine_coord(x, mat, coord_tfm, sz, mode, pad_mode, align_corners)
389 coords = affine_grid(mat, x.shape[:2] + size, align_corners=align_corners)
390 if coord_tfm is not None: coords = coord_tfm(coords)
→ 391 return TensorImage(_grid_sample(x, coords, mode=mode, padding_mode=pad_mode, align_corners=align_corners))
File ~/mambaforge/lib/python3.10/site-packages/fastai/vision/augment.py:364, in _grid_sample(x, coords, mode, padding_mode, align_corners)
362 if d>1 and d>z:
363 x = F.interpolate(x, scale_factor=1/d, mode=‘area’, recompute_scale_factor=True)
→ 364 return F.grid_sample(x, coords, mode=mode, padding_mode=padding_mode, align_corners=align_corners)
File ~/mambaforge/lib/python3.10/site-packages/torch/nn/functional.py:4197, in grid_sample(input, grid, mode, padding_mode, align_corners)
4097 r""“Given an :attr:input and a flow-field :attr:grid, computes the
4098 output using :attr:input values and pixel locations from :attr:grid.
4099
(…)
4194 … _OpenCV: opencv/modules/imgproc/src/resize.cpp at f345ed564a06178670750bad59526cfa4033be55 · opencv/opencv · GitHub
4195 “””
4196 if has_torch_function_variadic(input, grid):
→ 4197 return handle_torch_function(
4198 grid_sample, (input, grid), input, grid, mode=mode, padding_mode=padding_mode, align_corners=align_corners
4199 )
4200 if mode != “bilinear” and mode != “nearest” and mode != “bicubic”:
4201 raise ValueError(
4202 "nn.functional.grid_sample(): expected mode to be "
4203 “‘bilinear’, ‘nearest’ or ‘bicubic’, but got: ‘{}’”.format(mode)
4204 )
File ~/mambaforge/lib/python3.10/site-packages/torch/overrides.py:1534, in handle_torch_function(public_api, relevant_args, *args, **kwargs)
1528 warnings.warn("Defining your __torch_function__ as a plain method is deprecated and " 1529 "will be an error in future, please define it as a classmethod.", 1530 DeprecationWarning) 1532 # Use public_apiinstead ofimplementation` so torch_function
1533 # implementations can do equality/identity comparisons.
→ 1534 result = torch_func_method(public_api, types, args, kwargs)
1536 if result is not NotImplemented:
1537 return result
File ~/mambaforge/lib/python3.10/site-packages/fastai/torch_core.py:378, in TensorBase.torch_function(cls, func, types, args, kwargs)
376 if cls.debug and func.name not in (‘str’,‘repr’): print(func, types, args, kwargs)
377 if _torch_handled(args, cls._opt, func): types = (torch.Tensor,)
→ 378 res = super().torch_function(func, types, args, ifnone(kwargs, {}))
379 dict_objs = _find_args(args) if args else _find_args(list(kwargs.values()))
380 if issubclass(type(res),TensorBase) and dict_objs: res.set_meta(dict_objs[0],as_copy=True)
File ~/mambaforge/lib/python3.10/site-packages/torch/_tensor.py:1278, in Tensor.torch_function(cls, func, types, args, kwargs)
1275 return NotImplemented
1277 with _C.DisableTorchFunction():
→ 1278 ret = func(*args, **kwargs)
1279 if func in get_default_nowrap_functions():
1280 return ret
File ~/mambaforge/lib/python3.10/site-packages/torch/nn/functional.py:4235, in grid_sample(input, grid, mode, padding_mode, align_corners)
4227 warnings.warn(
4228 "Default grid_sample and affine_grid behavior has changed "
4229 "to align_corners=False since 1.3.0. Please specify "
4230 "align_corners=True if the old behavior is desired. "
4231 “See the documentation of grid_sample for details.”
4232 )
4233 align_corners = False
→ 4235 return torch.grid_sampler(input, grid, mode_enum, padding_mode_enum, align_corners)
RuntimeError: grid_sampler(): expected grid to have size 1 in last dimension, but got grid with sizes [3, 90, 160, 2]