The fastai library is incredibly helpful, but obviously it can’t be used for every scenario. I’ve run into situations where it seems either impossible or overly convoluted to use fastai for that particular use-case (sample-wise loss and multimodal models are two examples I’m currently struggling with). However, when trying to move away from fastai I fail to replicate fastai results. The point of this thread is to go through how to manually implement/replicate fastai results without fastai. I think this will give a lot of users a greater depth as to what is going on with fastai and allow us all to be more flexible with the principles we learn from fastai.
This first post will be asking for specific help on one step of this process of manually replicating fastai results. My goal is to continue to post on this thread as I move throughout the process. I’ll use the multimodal problem I’m working on as the use-case.
I have been using a fastai TabularLearner on just a tabular data set (all continuous features) and getting respectable results (about equivalent to my results using a random forest model) - this fastai TabularLearner is my baseline. I then tried combining this in a multimodal model with associated text data and got poor results. To debug, I went back to only tabular data but did not use fastai - got poor results. I double and triple checked my normalization technique, fit function, etc.; they all seemed good. I then tried a ridiculous thing, the results of which are baffling to me. I used fastai and left everything the same as my baseline, except I manually inserted the model:
import fastai.tabular as tab
import fastai.layers as fast_layers
# Make databunch
procs = [tab.Normalize]
train_val_data_df = d_in.loc[train_val_df_indices].merge(d_out.loc[train_val_df_indices][clf_target], left_index=True, right_index=True)
fast_data = tab.TabularDataBunch.from_df('.', train_val_data_df, clf_target, \
valid_idx=val_iloc_indices, test_df=full_test_df[in_cols], procs=procs)
# Model architecture
layers = [337, 290, 82]
dropouts = [0.3, 0.3, 0.07]
learner = tab.tabular_learner(fast_data, layers=layers, ps=dropouts, metrics=tab.accuracy)
weights = torch.Tensor([0.413250185, 0.586749815]).cuda()
learner.loss_func = fast_layers.CrossEntropyFlat(weight=weights)
# Added the line below and that's the only change I made
learner.model = tab.TabularModel(emb_szs=[], n_cont=len(in_cols), out_sz=2, layers=layers, ps=dropouts).cuda()
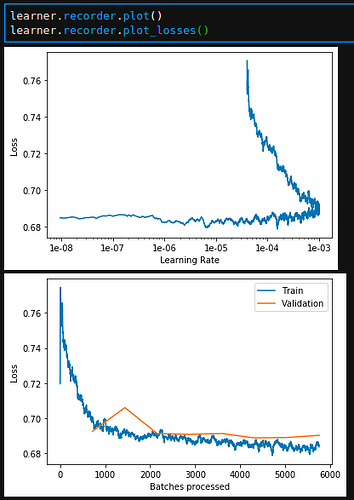
This model should be the exact same as the one that is created in tab.tabular_learner(...), but yet I get poor results when doing this. The losses look worse, as do my domain-specific results.
Baseline loss:
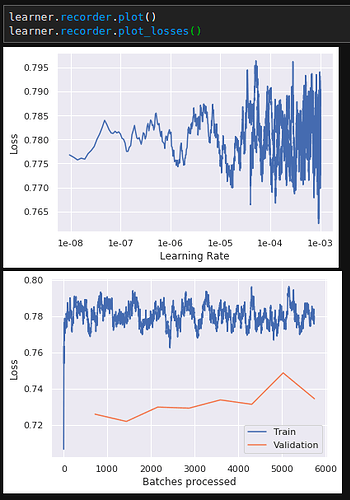
Manually inserted model loss:
Also the results of executing
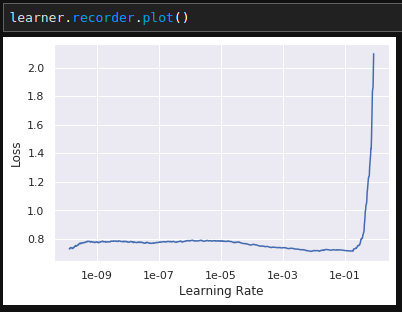
learner.lr_find(1e-10, 1, num_it=1000) also changes drastically.Baseline lr_find:
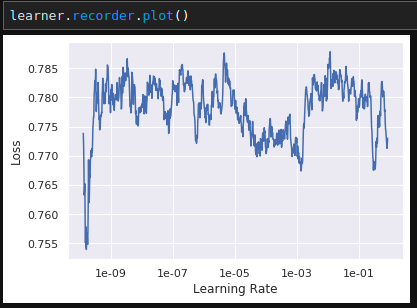
Manually inserted model lr_find:
The domain-specific metrics were substantially worse.
It seems to me like something must be happening in fastai’s tabular_learner that messes with the model after it instantiates it, but looking through the source code I can’t seem to find what it is. Any suggestions? What fastai magic am I not replicating? Recall that the only thing I changed was inserting manually the model that TabularLearner makes; I still used the same databunch and learner.fit functions as my baseline.