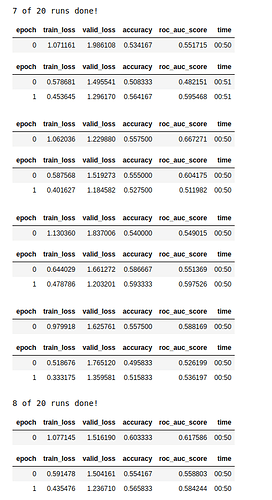
Hi, I am trying to do a gene prediction from pathology whole slide images. The results at the moment are far from good. (It’s not clear whether this gene is well predictable at all, nontheless I try what I can  ).
).
I was curious what more experienced deep learning people than me (i am a medical student) would spontaneously think when they would see these “bumpy” results? - I don’t know what to do against the overfitting for example in several runs.
(I also sometimes have results a little better than whats shown in the image, but I don’t expect to be better than a 60% accuracy averaged per medical case. (accuracy and auroc here are shown per tile, after all runs I also average for accuracy and auroc per case which are usually about +2 percent better than per tile, I use 50 tiles per patient.)
Some more information:
I do a 4-fold cross validation (random undersampling before) with 50 tiles from 96 medical cases each (each time there are 24 cases = 1200 tiles in the validation set). It’s a binary classification. I put different kinds of mutation from the same gene together and then made something like “clinically relevant are a yes”, non-clear ones are excluded and the other cases get the “negative” label.
I am using a pytorch pretrained efficientnet-b2a, batch size = 55 (tiles are 512*512 pixels) and a weight decay of 0.1 and I do 2 epochs of “fine_tune” for every split (no learning rate specified).
I use a little image augmentation too (random flipping and some little color disturbations).
I do fine tuning with every fold after each other (so 1 run equals once through all 4 folds). the model gets created new each time before a new split.
At the end I average the results over for example 20 runs here.
Any feedback highly appreciated