That sounds very cool. I’m not sure it’ll work in practice, but let’s ask: @nolliag @matt.mcclean is this something that can be integrated in to the lesson 1 notebook?
Exciting news in the Sagemaker world - you can now create larger volumes for your work! 
@matt.mcclean would you be able to update your script? I’m thinking it should work like this:
- Install everything as now, if not already done
- Create a 50GB volume
- Add an empty executable shell script in the persistent volume
Then on every run do the following:
- Symlink .bashrc, .fastai, and anaconda3 from persistent volume to homedir
- Run the (initially empty) executable shell script added earlier
That way, the user can fully customize their anaconda install, and their shell, etc, without needing to change the cloudformation script.
How does that sound?
cc @nolliag
Yes, great news for sure  will update it now
will update it now
Also, need to fix the image links in the SageMaker docs as they are broken
@avishalom it depends. If you want rapid iteration of training your fastai models then it would be best to launch a p2.xlarge or p3.2xlarge SageMaker notebook instance type. You have a dedicated GPU which can be used at any time.
You can use a t2 Notebook instance type to develop your code and use the p2 or p3 for the SageMaker training service however it will add a few extra minutes for SageMaker to provision the instance where the training will be executed. The SageMaker training service uses Docker images when training models so you will need to provide a custom Docker image with fastai libraries installed and uploaded to your Elastic Container Registry (ECR). I have an example Github project here showing how you can use the SageMaker training and hosting service for your fastai models.
@avishalom what I did is added a script which is being shared by you while creating the notebook but I am getting the same error on starting the notebook again.
So do I need to manually type the code below on terminal everytime I start the notebook :
cd /home/ec2-user/SageMaker
source activate envs/fastai
ipython kernel install --name ‘fastai’ --display-name ‘Python 3’ --user
Hi,
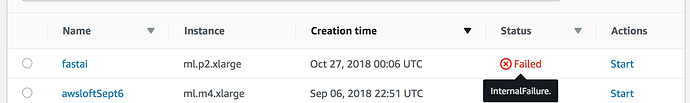
I have been working on sagemaker for past few days, and everything was fine. I tried to start the notebook instance now and it failed. The error message is not helpful.
It says Internal Failure. Here is the screenshot.
Since I tried multiple datasets, I think its something to do with less memory.
Did I lose the work that I did so far? Should I create new instance and start fresh?
This isn’t a good place to ask for individual account support. Please use official AWS support channels for that.
Thanks for @matt.mcclean we now have an updated installation experience which should “just work”. All your data and models will be stored persistently, and you’ll have plenty of space.
https://course-v3.fast.ai/start_sagemaker.html
Please try it out (delete your old config and instances first) and let us know how it goes for you!
Did you try and have a look at the CloudWatch logs to see what the error is when restarting the instance? You can goto the CloudWatch console by clicking the Services option on your AWS console then typing in cloudwatch to the search box. When your CloudWatch page is open click the Logs option in the left sidebar menu and look for the Log Group named /aws/sagemaker/NotebookInstances. Click on it then select the Log Stream named fastai/LifecycleConfigOnStart. You should hopefully see the error message why it did not work. You should not have lost the work so far as it is saved to the EBS volume so persisted between restarts. If you create a new instance you will lose your previous work.
Thanks Matt . I could not find any helpful log (no error message). But I ended up creating new instance since I could not open the old one. I convinced myself that I lost the work! Thankfully this happened after 1 week, and not towards the end of the course.
If it happens again then I would delete the Start notebook script commands so it should boot up fine. That way you will not lose your previous work. You will have to run manually the following commands in a terminal in your Jupyter web console to create the symlinks and install the ipython kernel.
ln -s /home/ec2-user/SageMaker/.torch /home/ec2-user/.torch
ln -s /home/ec2-user/SageMaker/.fastai /home/ec2-user/.fastai
cd /home/ec2-user/SageMaker
source activate envs/fastai
ipython kernel install --name 'fastai' --display-name 'Python 3' --user
@matt.mcclean Finally what we need to follow for installation ? Do we need to delete the notebooks which we had created earlier using the official fastai documentation and follow the new one to install?
I couldn’t find the Launch Stack Button. Can anyone help me with step 2 and 3 of Sagemaker setup?
The Launch Stack button is found on this page
The “Launch stack” buttons are on this page, https://course-v3.fast.ai/start_sagemaker.html#getting-set-up, not your aws console. They look like they are part of the guide photos, but are actual buttons.
I am assuming that the new script by matt solved it ?
sorry, i was away for a bit.
@ everyone else, is there an easy way to add an “at now + 4 hours shutdown” or something at the start of the notebook, to prevent people forgetting to turn it off.
also, when returning to a notebook that was created earlier, how do i get the content to update (new ipynb notebooks are added weekly, but do not appear on the created instance)
FWIW the stack method isn’t working for me, “course-v3” folder is not appearing 20 minutes after cloudwatch seems to have completed creating the notebook (@matt.mcclean )
At the moment you cannot automatically stop a SageMaker instance. You could do this with a Lambda function that is triggered by a CloudWatch event, every night for example at a certain time to check if your notebook is running and stop it if so. Here is a link to the AWS documentation showing how to run a Lambda function on a schedule.
If you have launched a SageMaker notebook recently using the CFN template then it will install a custom start script in the directory ~/SageMaker/custom-start-script.sh. You can modify this to perform custom actions when the notebook is restarted. The default behaviour is to update the fastai conda library and also update the course-v3 repo from GitHub.
Regarding the error, did you have a look in the CloudWatch log file as per the instructions here?
can’t we just stop our notebook instances by just clicking the stop button as suggested in official doc??