I currently have nbdev 2.2.6, which I got by running pip3 install --upgrade --force-reinstall nbdev.
It’s only in master at present. Probably will do a release with it today.
Thank you Wasim, this is helpful.
I just updated to the latest nbdev and did get the warnings now. However, since I have got lots of mixes in more than one notebooks, and the warning does not tell me where those mixes come from, I wonder whether you could add the info on where those mixes are from or much better if nbdev could automatically separate the import into a separate cell?
FYI @Daniel I’m tracking this as an issue here Show location of mixing of imports and code · Issue #938 · fastai/nbdev · GitHub. Thank you for suggesting this
Great, thanks! nbdev is amazing and I am using it everyday.
I am now using nbdev daily, and I am very impressed by how good it is. The only point I miss (which is not nbdev’s fault) is to turn jupyter lab into a more IDE-like experience.
I know this might be the wrong venue since this is not directly related to nbdev, but I would like to know how nbdev users deals with this? I miss features like code refactoring, searching through object names, jumping back and forth through definitions, etc.
After a quick search, I found jupyterlab-lsp but haven’t tried it yet. Any suggestion?
@Daniel Your enthusiasm, and the care and curiosity that you put into writing your posts, make me very happy to be working on nbdev! ![]()
I do miss this too btw. I sometimes go back to a plaintext editor for hopping around and reading code
Otherwise, the options I know of are:
- VSCode’s notebook editor has builtin LSP support
- JetBrain’s PyCharm or DataSpell have something similar, I think
- I’ve seen people recommend the Lab LSP plugin you mentioned but haven’t tried it myself
It seems that nbdev_test runs the notebooks in parallel. This is convenient for speed but can be tricky when running CI.
For example, one of my notebooks creates a file on disk and removes it at the end. Another notebook creates the same file but first checks to make sure the file does not exist before creating it. Since both notebooks are running in parallel the file created on notebook 1 makes the test fail in notebook 2.
jobs:
test:
runs-on: ubuntu-latest
steps: [uses: fastai/workflows/nbdev-ci@master]
Is it possible to control which notebooks are run in parallel and which are run sequentially when using fastai/workflows/nbdev-ci@master?
I think for something more complicated like that, you might be better off copying the underlying workflow and editing it to suit your needs. You can set the number of workers to 0 to disable parallel tests
Personally, I would try to find a way to make the tests parallel-friendly. Could they use different files during testing?
Some code refactoring techniques don’t work well with nbdev, although nowadays I rarely want auto-refactoring since in a notebook-first process I do all the exploration interactively and generally get to a pretty good solution that way. For stuff like symbol renaming I generally just use perl -pi -e 's/.../.../' file.ipynb.
As @seem says though, jumping over to an IDE/editor from time to time can be useful. Especially for jumping around a code-base you’re not familiar with. And of course, you can – since all the .py files are there and can be treated in the usual way. (I don’t even do that as much as I used to, since I use ?? in jupyter a lot and that’s often all I need.)
Many refactorings done in an editor can be synced back to the notebooks using nbdev_update BTW.
Yes, my first thought was to copy and modify the workflow file, which is what I will do. The disadvantage is that I will lose automatic updates that you guys eventually do in the workflow.
About having different file names, that was my second thought, but the issue here is that notebook2 is a sequence to notebook1 and uses the same file as in notebook1, so changing the name would help the CI but would make following the tutorial a little more confusing.
Yes, I haven’t used nbdev_update yet, but it seems useful for code refactoring.
One basic need I have is when my notebook gets big, and I want to open it and go straight to the point where I want to modify it. I use the find functionality, but that search over text is slow compared to IDE, which filters only function names, for example.
I will give it a try to VS, as @seem suggested, as it is something I want to try anyway, as they seem to be making a lot of progress on ML/AI productivity.
Can you explain more about what you mean here?
Sure. Currently, my test.yaml looks like this;
jobs:
test:
runs-on: ubuntu-latest
steps: [uses: fastai/workflows/nbdev-ci@master]
I understand that if the maintainer of fastai/workflows/nbdev-ci@master make code changes to this workflow, it will automatically propagate to my CI.
If I copy the contents of the workflow above to my repo and modify it to fix the issues I mentioned, I will no longer be in sync with future changes made in fastai/workflows/nbdev-ci@master.
Yes that’s true. I use the Collapsible Headings and TOC2 extensions to move around notebooks more quickly, but it’s not as fast to jump to a definition as with a regular editor.
Failed to clean notebooks and URI malformed
I have written a library for a second time and this time I was fully consumed by it and didn’t run nbdev_prepare until now. After fixed all the mix cell issues, I still got two issues (I did update nbdev though):
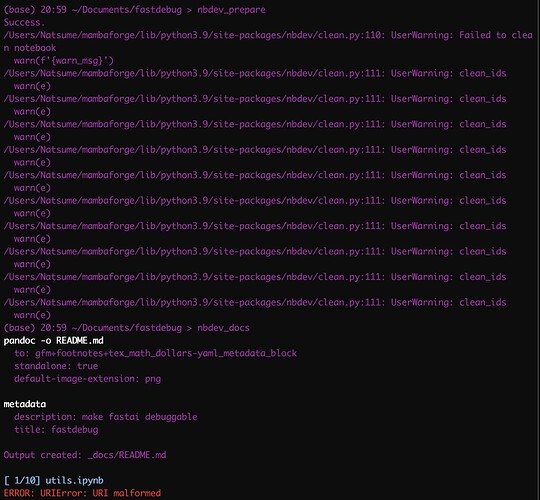
The first issue is after I run nbdev_prepare I got a lots of messages saying 'failed to clean notebook"
The second issue is after I run nbdev_docs, I got “URI malformed”
Could anyone help me have a look? Thanks
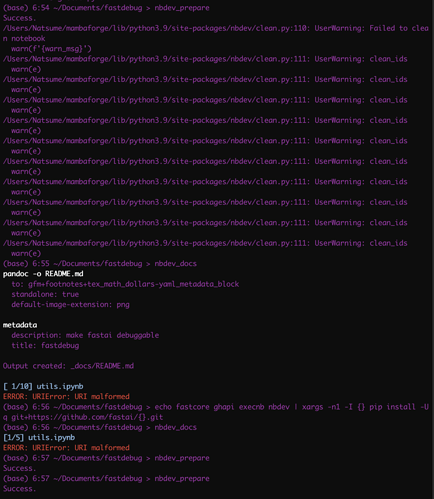
Thanks a lot @tgmstat, and the solution by updating to the latest libraries with git by @hamelsmu solved the first issue for me. However, I still have no idea where to look to fix my second issue with 'URI malformed"
Can you tell me more about the URI malformed? My apologies as I may have missed that or lost track of that one!