This might be a really stupid question, but when you pass in a metric, such as RMSE, how is it being calculated during training?
During training I am getting pretty good results, but then when I call .predict() on the model (loaded via best_save_name), and manually calculate the RMSE using the same function, I am getting a completely different number (and I’m using the same validation indices for y_true).
Also, when validating (at the end of each epoch during training), it iterates through 1236 validation samples which, with a batch size of 128, accounts for only 17.5% of my data (and is only half of my validation size, which is 35%).
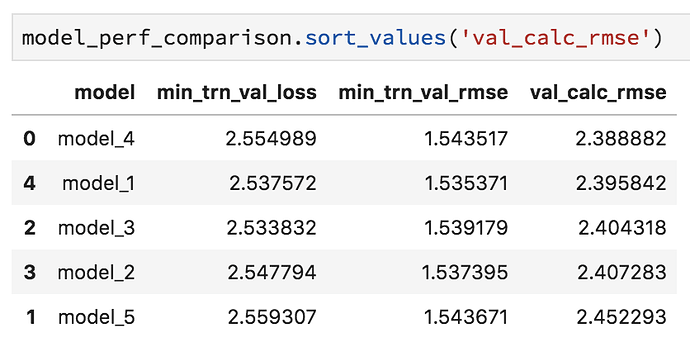
Here is a screenshot of the min loss and rmse shown during training vs the rmse calculated with the best save:
As you can see, there isn’t even a relationship between the best performance…
So I really have no idea how this is being calculated during training! I really hope someone can clarify what is probably a simple misunderstanding.
Thank you!!
Edit: and just to make sure there is no bug in how I’m calculating the loss of the saved models, here is the function:
def load_model_get_val_rmse(saved_weights, loc_val_idx):
# init model objects
md = ColumnarModelData.from_data_frame(path = 'models', val_idxs = loc_val_idx, df = df,
y = yl.astype(np.float32), cat_flds = cat_vars,
bs = 128, test_df = df_test)
m = md.get_learner(emb_szs = emb_szs, n_cont = len(df.columns) - len(cat_vars),
emb_drop = 0.04, out_sz = 1, szs = arch, drops = dropout, y_range = y_range)
# load saved weights
m.load(saved_weights)
# calc rmse
yl_true = deepcopy(yl[loc_val_idx])
yl_pred = deepcopy(m.predict().reshape(-1,))
error = rmse(yl_pred, yl_true)
return error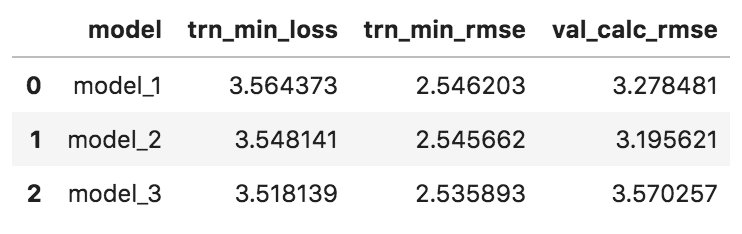
 )… I’ll test this out with a few models and get back to you to confirm if this helps
)… I’ll test this out with a few models and get back to you to confirm if this helps