thanks for this post. pip install dtreeviz resolved my issue.
In the TabularPandas and TabularProc section of 09_tabular.ipynb
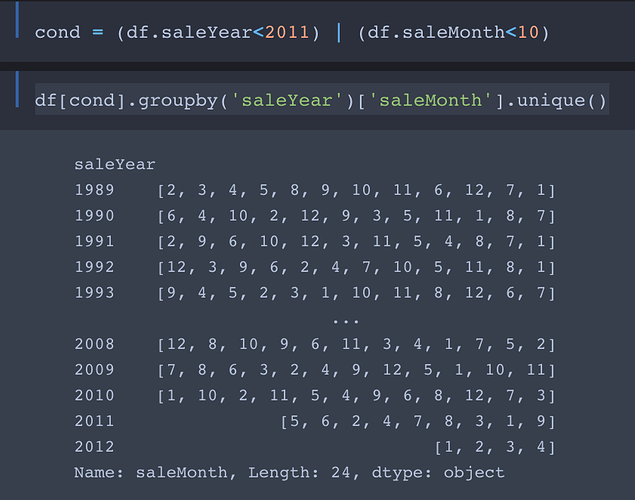
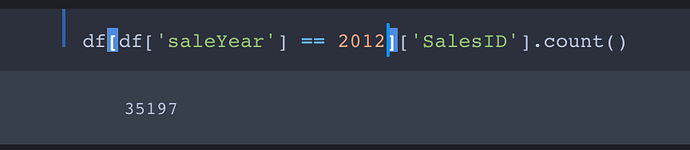
We are splitting the training and the validation set with before November 2011 and after November 2011
cond = (df.saleYear<2011) | (df.saleMonth<10)
train_idx = np.where( cond)[0]
valid_idx = np.where(~cond)[0]
splits = (list(train_idx),list(valid_idx))
Should the logic be as follow ?
cond = (df.saleYear<2011) | ( (df.saleMonth<10) & (df.saleYear==2011) )
In Chapter 8 Collab, the embedding can be created with
def create_params(size):
return nn.Parameter(torch.zeros(*size).normal_(0, .01))
Is the std equal to 0.01 because n_factors is 50, and so 1/50 \sim 0.01?
On a related note (and apologies if I’ve misread the text), in this paragraph
To calculate the result for a particular movie and user a combination we have to look up the index of the movie in our movie latent factors matrix, and the index of the user in our user latent factors matrix, and then we can do our dot product between the two latent factor vectors. But look up in an index is not an operation which our deep learning models know how to do.
it seems to say that we cannot simply index into an embedding, but doesn’t this implementation later on in the chapter
class DotProductBias(Module):
def __init__(self, n_users, n_movies, n_factors, y_range=(0,5.5)):
self.user_factors = create_params([n_users, n_factors])
self.user_bias = create_params([n_users])
self.movie_factors = create_params([n_movies, n_factors])
self.movie_bias = create_params([n_movies])
self.y_range = y_range
def forward(self, x):
users = self.user_factors[x[:,0]]
movies = self.movie_factors[x[:,1]]
res = (users*movies).sum(dim=1)
res += self.user_bias[x[:,0]] + self.movie_bias[x[:,1]]
return sigmoid_range(res, *self.y_range)
show that yes we can? Aren’t the square brackets on user_factors and movie_factors the same as ‘look up in an index’?
I’m working in paperspace and am getting the same errors. I had already imported utils from fastbook. Not sure why
NameError: name ‘draw_tree’ is not defined
Hi @Grace1 Try this: make sure that you
cd into the fastbook folder
before you import utils.
Here is what I did:
%cd '/content/drive/My Drive/fastbook/'
from utils import *
%cd ..
For reference, please see my Colab notebook.
Yes, I also think that it should be the AND operator, &.
200_000 == 200000. Nice.
If you don’t use small NNs, the combined size of your model gets large very fast. The question then is if you shouldn’t just use a single large NN instead.
What’s “entity encoding”? I don’t see the term in the linked article.
Thought that was a type and corrected it!
No, the use of | is actually intended. Here’s why:
We want the training set to contain all entries with saleYear up to September 2011 and the validation set to contain all entries from October 2011 forward.
So the first part of cond selects entries with saleYear of 2010 and earlier into the training set. The second part filters the entries with saleYear ranging from from Jan 2011 to Sept 2011 into the training set.
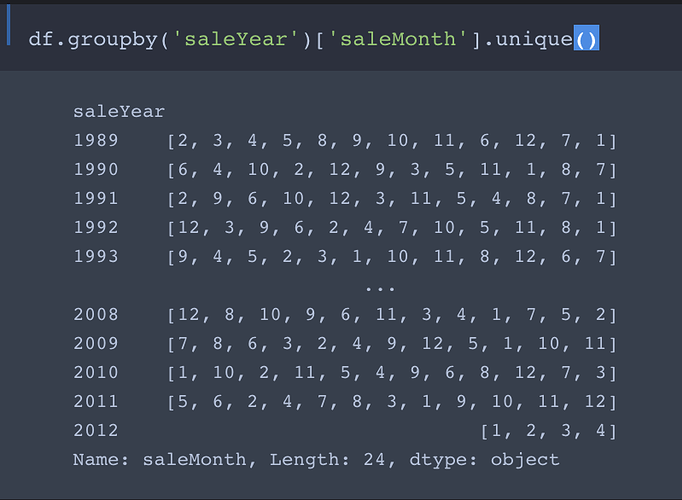
Comment: In my opinion this not a good way to do the filtering. As you may have noticed, it works only if there are no entries with saleYear later than 2011. If we were to add entries from the first three quarters of 2012 (or any later year) to the data, they would get (unintentionally) thrown into the training set!
A better, foolproof way to filter the data into the training and test sets is
cond = (df.saleYear<2011) | ( (df.saleMonth<10) & (df.saleYear==2011) ))
Great catch @sylvaint!
With the corrected filtering algorithm, we should be able to get improved accuracy on the predicted sale prices!
Actually it does not help. Having more recent data in the training set skewed the results positively.
With the correct splits I get:
| Model | Updated r_mse | Previous r_mse |
|---|---|---|
| random forest | 0.24241 | 0.230631 |
| neural net | 0.263618 | 0.2258 |
| ensembling | 0.241996 | 0.22291 |
The results are not as good as with the erroneous splits. More importantly the neural net result is not as good as the random forest result which contradicts the book’s section conclusion for that dataset.
Well done, @sylvaint!
I was going to change my previous comment to predict that things would in fact get worse rather than better, but I see I’m too late.
So, I’ll explain why things got worse:
Performance on the corrected validation set becomes poorer than with the original validation set because of data leakage, which Jeremy also discusses in Chapter 9. Indeed, this case serves as an excellent example.
Before we corrected the filter for the training and validation sets, the model was able to (unintentionally) cheat by data snooping, i.e. by illegally incorporating into the training set some of the data from the later times (post-2010) which it is trying to predict. This is an example of data leakage So with the original training and validation split, the validation scores are artificially high due to cheating.
The corrected filter purges the training set of the illegal data points, which now become part of the validation set, as they should have been in the first place. Now, not only can’t the model cheat, but it has to predict on even more post-2010 data points, which is why the validation scores become worse!
What is the best way to extract the embeddings from the tabular_learner or TabularModel object?
If you want the weights you should be able to just do learn.model.embeds.weights