The video, notebooks, spreadsheets, and links are available here.
Embedding - In lesson4, under the Dot Product section, Embedding is called to try to create an embedding layer.
u = Embedding(n_users, n_factors, input_length=1, W_regularizer=l2(1e-5))(user_in)
It gives me a NameError and I am not sure how to fix it. I don’t know if it is a setup problem on my side (I set up my own server) or where I should be looking to fix this problem. I did a %whos and Embedding is not in my environment. Any suggestions on where I should go from here …
-=-=-
user_in = Input(shape=(1,), dtype=‘int64’, name=‘user_in’)
u = Embedding(n_users, n_factors, input_length=1, W_regularizer=l2(1e-5))(user_in)
movie_in = Input(shape=(1,), dtype=‘int64’, name=‘movie_in’)
m = Embedding(n_movies, n_factors, input_length=1, W_regularizer=l2(1e-5))(movie_in)
NameError Traceback (most recent call last)
in ()
1 user_in = Input(shape=(1,), dtype=‘int64’, name=‘user_in’)
----> 2 u = Embedding(n_users, n_factors, input_length=1, W_regularizer=l2(1e-5))(user_in)
3 movie_in = Input(shape=(1,), dtype=‘int64’, name=‘movie_in’)
4 m = Embedding(n_movies, n_factors, input_length=1, W_regularizer=l2(1e-5))(movie_in)
NameError: name ‘Embedding’ is not defined
-=-=-
%whos
Variable Type Data/Info
Adam type <class ‘keras.optimizers.Adam’>
BatchNormalization type <class ‘keras.layers.norm<…>tion.BatchNormalization’>
Convolution2D type <class ‘keras.layers.conv<…>olutional.Convolution2D’>
Dense type <class ‘keras.layers.core.Dense’>
Dropout type <class ‘keras.layers.core.Dropout’>
Flatten type <class ‘keras.layers.core.Flatten’>
GlobalAveragePooling2D type <class ‘keras.layers.pool<…>.GlobalAveragePooling2D’>
Image module <module ‘PIL.Image’ from <…>-packages/PIL/Image.pyc’>
Input function <function Input at 0x7f7aba5652a8>
K module <module ‘keras.backend’ f<…>as/backend/init.pyc’>
.
.
.
Thanks for the question. According to the docs, the full name is “keras.layers.embeddings.Embedding”. Therefore you can either refer to it by its full name everywhere you use it (which you probably don’t want to do!), or you can add to the top of your notebook:
from keras.layers.embeddings import Embedding
Or you can add that line to utils.py and reload it.
Here’s some more information about how python handles this.
Thanks, Jeremy. That worked. Glad its not a sign of something inherently wrong with my setup.
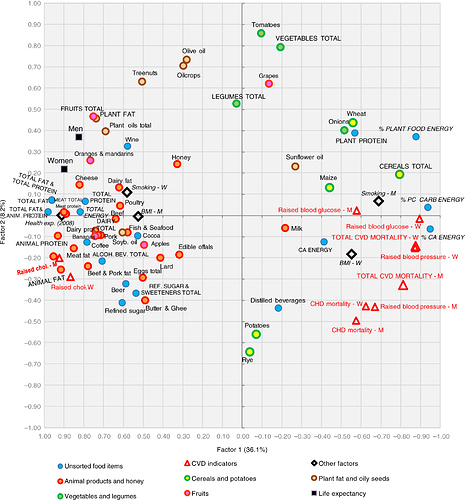
Lesson 4 reminded me of a “Factor analysis” I saw in a recent paper:
Source: http://www.foodandnutritionresearch.net/index.php/fnr/article/view/31694
Is this not very similar to the Koren et al chart @rachel posted in Slack?
Somewhat similar, yes. Factor analysis / principal components analysis are linear methods that create a lower dimensional matrix that attempts to capture the variance in the original matrix, just like our model did. If you want to read more, search for ‘PCA’ and ‘factor analysis’ for the classic methods like you show, and ‘probabilistic matrix factorization’ for the approach we used in class.
Very helpful, thank you!
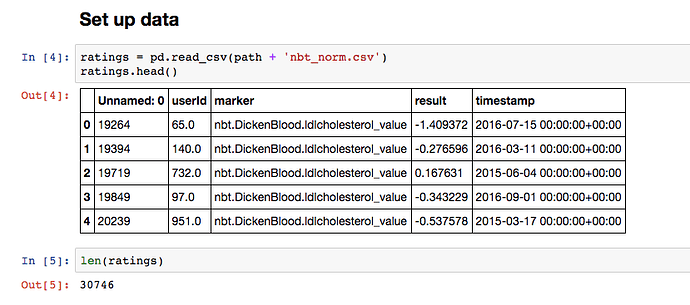
I tried the lesson 4 notebook using my own data:
30,746 total features taken from my database of blood tests, urinary organic acids and hormone metabolites. 447 unique users and 208 unique markers. I normalised everything using sklearn.preprocessing.scale.
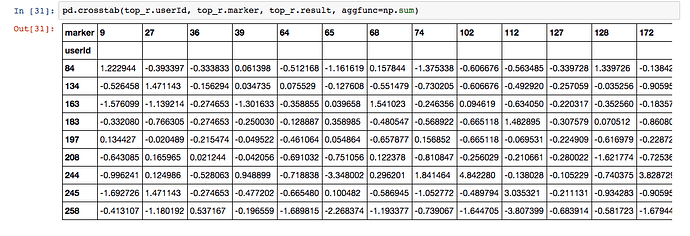
I reduced the number of latent factors to 5 and he rest of the notebook is as you presented it.
val_loss: 0.9314
I’m very excited about the idea of being able to describe my users with just five latent factors. I’m even more excited about scatter plotting the markers using their top latent factors because I think the plot will make physiological sense in the same way as the scatter plot for movies makes sense.
My question is, where did the latent factors in the Keras model go?
model.fit([trn.userId, trn.marker], trn.result, batch_size=64, nb_epoch=10,
validation_data=([val.userId, val.marker], val.result))
trn.result.shape
(24466,)
The users and markers were concatenated to form the input, do I need to split them back up to get my latent factors?
We’ll see how to access the latent factors tomorrow 
Huh, Google and ye shall find:
Warning - some of the code in that project is not too nice…!
Did you notice who he is? “Chief Architect at Elsevier, the world’s leading scientific publisher.”
@jeremy By the method of collaborative filtering, we need to predict the missing ratings right?
In the spreadsheet case: MovieId : 49 UserId: 212
result is 0.0 in the spreadsheet. Are we not supposed to predict this unknown rating, am I missing anything here ?
At training time (which is shown in the spreadsheets) we predict the rating for those which are labeled, so that we can compare to the true labels and calculate the value of the loss function. So we don’t predict the rating you mention, since we don’t have a label for it in the training set.
Just curious, how do we predict MovieId : 49 UserId: 212 value at test time ?
At this point: https://youtu.be/V2h3IOBDvrA?t=10m56s did @jeremy by any chance mean sigmoid is the activation function as opposed to loss function?
Yes! Thanks for correcting my silly mistake 
@jeremy I was able to solve the book-crossing dataset recommendation using similar architecture as taught in the class.
Below are results on a sample of 60k data points. Need to tune still since val loss is increasing…
Train on 48036 samples, validate on 11962 samples
Epoch 1/5
48036/48036 [==============================] - 34s - loss: 14.5920 - val_loss: 12.4964
Epoch 2/5
48036/48036 [==============================] - 46s - loss: 8.5997 - val_loss: 13.8392
Epoch 3/5
48036/48036 [==============================] - 78s - loss: 5.2262 - val_loss: 14.6633
Epoch 4/5
48036/48036 [==============================] - 78s - loss: 4.3011 - val_loss: 14.7733
Epoch 5/5
48036/48036 [==============================] - 79s - loss: 3.8658 - val_loss: 14.7772
Out[19]:
Try adding l2 regularization.
Hi All,
My buddy pointed me to this course a couple weeks ago, and I’m now totally obsessed with deep learning & neural networks. Thanks to everybody working on this course!
Anyway, I’m working through the homework, and I have a practical question about the embeddings: once a model is trained and set up, how do you add new users and new movies to the database?
Since the embeddings take as an input the number of users and number of movies as inputs (and create parameters accordingly), it seems the model is locked down to those movies and users.
Are there clever ways to modify the layers to include extra movies and users, or do you need to retrain from scratch every time you add new data?
Thanks,
-Caleb