I finally came across a solution for this here. I reloaded my data paths, created a learn object which allowed me to load the previously saved weights.
I trained “food 101” on Google Cloud Platform (Tesla P4 GPU). Took like ~7,5 min/epoch.
Hi, I am struggling about how to upload and prepare my dataset. I tried a few different ways but encountered different problems. Can anyone tell me how to solve these?
Q1) When I use another link from fastai dataset, an attribute error showed up. I tried untar_data(url=‘http…food-101.tgz’) but another error (not a gzip file) appeared.
SOLVED by removing .tgz at the back , referring to untar-data-requires-tgz-file-ending
https://colab.research.google.com/drive/1EFZDzaOECuZ23WPbJkBxzLuTh3Tbj-ov#scrollTo=BBHv5M-AKjtw
FOODS=‘https://s3.amazonaws.com/fast-ai-imageclas/food-101.tgz’
path = untar_data(URLs.FOODS)
path
AttributeError Traceback (most recent call last)
<ipython-input-4-1bee2d7d1f5a> in <module>() 1 FOODS=‘https://s3.amazonaws.com/fast-ai-imageclas/food-101.tgz’ ----> 2 path = untar_data(URLs.FOODS) 3 path
AttributeError: type object ‘URLs’ has no attribute ‘FOODS’
Q2) I prepared my own dataset with google image download as suggested in another thread. But i had a hard time uploading it to Colab. After uploading the file to google drive, I tried the code snippet in colab to import data from google drive. I ended up with a GoogleDriveFile which is not directly usable. What should I do next?
I used path = Path(root_dir + ‘Colab Notebooks/CatCategoryDS’) to solve the problem. But i would love to know if anyone know how to handle GoogleDriveFile. Thanks!
Q3) I feel like I need a lot of practice and learning in data preparation and handling. But I cant find much resources online. Any recommendations??
Thank you so much for the advice!
I have training data in an NPY file whose shape is (814, 440, 440, 1), which consist of 814 images of size 440 x 440 x 1, and the label is an (814, 1) NPY file containing ones and zeroes.
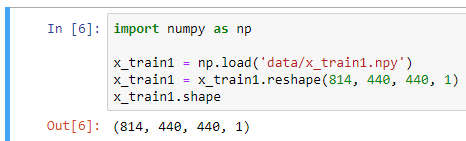
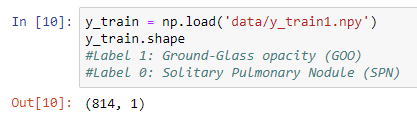
I used to train this with Keras, using model.fit
![]()
Is there a way to create a data bunch with the fastai library from these data?
Thank you in advance!
Hello! I am new to lesson 1.
My configuration is Win10+anaconda+python3.7+torch1.0.1+fastai1.0.50.post1+cuda80
After I trained ResNet34 successfully, i try to continue with ResNet50.
By checking the nvidia-smi, the memory is not used:
+-----------------------------------------------------------------------------+
| NVIDIA-SMI 398.35 Driver Version: 398.35 |
|-------------------------------+----------------------+----------------------+
| GPU Name TCC/WDDM | Bus-Id Disp.A | Volatile Uncorr. ECC |
| Fan Temp Perf Pwr:Usage/Cap| Memory-Usage | GPU-Util Compute M. |
|===============================+======================+======================|
| 0 GeForce GTX 960M WDDM | 00000000:01:00.0 Off | N/A |
| N/A 47C P8 N/A / N/A | 40MiB / 4096MiB | 0% Default |
+-------------------------------+----------------------+----------------------+
+-----------------------------------------------------------------------------+
| Processes: GPU Memory |
| GPU PID Type Process name Usage |
|=============================================================================|
| No running processes found |
+-----------------------------------------------------------------------------+
But I still came across the following CUDA out of memory
and
Tried to allocate 1024.00 KiB (GPU 0; 4.00 GiB total capacity; 3.08 GiB already allocated; 588.80 KiB free; 12.20 MiB cached)
I tried to reboot and it doesn’t work!
How can i do next? thanks in advance~
Hi, I’m having same issue, implementing a custom metrics about F1 score.
I have
len(data.train_ds) = 750,
len(data.valid_ds) = 250.
In my metrics function, I have at the bottom:
precision = TP/(P + 1e-12)
recall = TP/(CP + 1e-12)
print('prec: ', precision.shape)
print('recall: ', recall.shape)
F1 = 2 * (precision * recall) / (precision + recall + 1e-12)
return torch.Tensor(F1)
During fit_one_cycle() it prints:
prec: (12,)
recall: (12,)
prec: (12,)
recall: (12,)
prec: (11,)
recall: (11,)
And this is the ending part of the output:
/opt/conda/lib/python3.6/site-packages/fastai/callback.py in on_batch_end(self, last_output, last_target, **kwargs)
343 dist.all_reduce(val, op=dist.ReduceOp.SUM)
344 val /= self.world
–> 345 self.val += last_target[0].size(0) * val.detach().cpu()
346
347 def on_epoch_end(self, last_metrics, **kwargs):
RuntimeError: The size of tensor a (12) must match the size of tensor b (13) at non-singleton dimension 0
Any suggestion about how to solve it? ^^
Hi Yujie,
What batchsize are you using when training with resnet50? Resnet50 will take much more memory than Resnet34, so you should try halving the batchsize (compared to your resnet34 batchsize). For example, if in resnet34 you used batchsize 64, then try batchsize 32 for resnet50 and see how that goes.
Yijin
Hey guys. I just finished my first course and I have so many doubts. Firstly how do we know how to select the number of fit_one_cycle? I saw some replies saying we have to choose such that they dont overfit. Does it mean we have to experiment with few numbers and choose the one where the training and validation loss isnt much different??
We have been asked to pick an image classification problem post the lecture. I was thinking of trying out SVHN data. But I am not sure if I can solve that at the current stage of my knowledge. Should I pick a simpler problem and wait for few more lectures before trying to solve SVHN?
I’ve been trying to practice on kaggle datasets with some of the code from lesson 1, however, I’m running into an issue with generating predictions on a test set. I tried learn.get_preds(is_test=True), which did not work, and also tried looping through the images one by one. The last method works. but it takes 40 seconds for each prediction. Is this normal? With 4000 images to predict, it seems unusually long.
You are right about that. Try with few cycles/epochs and stop when you see your model overfit.
I can see from the output of Resnet50 that the model is confused between Egyptian Mau and Bengal cat, which really look the same except for the COLOR. One of these is really golden brown.
@jeremy- Do I need to add more layers to the model to handle color better or is there something else that you would recommend?
I’m having trouble understanding how Jupyter works with this course. So I opened up a Crestle account and then, from Crestle, opened the course Jupyter notebook. Once I’ve done that, can anyone else change the notebook? Can I make changes to it and have them saved? Is this Jupyter notebook linked to my Crestle account or is there another way to access it?
Also I’m sorry if I’m not replying to the right person, I’m not really sure how to use this forum.
Wasnt the maximum learning rate taken for session 1 because it had a higher loss associated to it? Like i used 1e-6 to 1e-4 and the loss is like 0.91. Then i changed it to 1e-1 and the error increased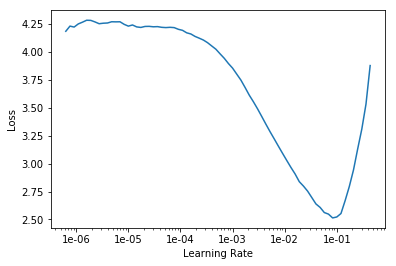
This is my learning rate graph. Kindly do help me fine tune it!
Hokie, I think crestle.ai team might be able to answer your questions re: the visibility of your notebooks to others. I presume you are trying to share your progress with someone by sharing a link to it?
AFAICT, since you have to log in to your account in Crestle, you’d be the only one who has direct access to your jupyter notebook.
Also, Jupyter automatically saves your work, at least on my local machine it does. I’m not sure what the behavior is for the hosted jupyter notebooks at crestle, I would assume that they also do this.
I would suggest you put these questions to Crestle support to get definitive answers.
One thing that Jeremy recommends in his first lecture is to make a copy of the notebook and make changes in that notebook (because jupyter saves your changes every few seconds/minutes)
I haven’t played with the new crestle.ai notebooks so I’m not sure how they behave. I’m assuming they behave like regular jupyter notebooks, except that they are hosted for you on a machine in the cloud and you can login to your area on their servers and run the notebook on their servers.
Is Jeremy’s guide to creating our own dataset up-to-date? I read through it and saw a lot of error/warning messages underneath the code chunks. Is that a problem? I don’t know whether to ignore that.
edit for example:
Error https://npn-ndfapda.netdna-ssl.com/original/2X/9/973877494e28bd274c535610ffa8e262f7dcd0f2.jpeg HTTPSConnectionPool(host=‘npn-ndfapda.netdna-ssl.com’, port=443): Max retries exceeded with url: /original/2X/9/973877494e28bd274c535610ffa8e262f7dcd0f2.jpeg (Caused by NewConnectionError(‘<urllib3.connection.VerifiedHTTPSConnection object at 0x7f2f7c168f60>: Failed to establish a new connection: [Errno -2] Name or service not known’))
Hey, I’ve been trying to import a kaggle dataset onto the kaggle kernel.
The folder set up is as follow:
/input
/train_images
/train.csv
the images are in folder train_images and labels are present in train.csv.
i tried to use ImageList.from_csv(path, ‘train.csv’, cols =2) but it looks for the label filename in input folder and not in train_images folder. However if i try to use ImageDataBunch.from_csv(path,ds_tfms=tfms, size=28) then the function looks for only ‘labels.csv’ specifically and returns an error saying no folder called ‘…/input/labels.csv’ because the file is called ‘train.csv’.
And i cannot rename the file either cause kaggle doesn’t allow writes onto data and hence I’m unable to rename.
error with line 17
doc(interp.plot_top_losses)
gives
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
<ipython-input-17-a7fe6964f6f7> in <module>
----> 1 doc(interp.plot_top_losses)
~/miniconda3/envs/p37cu10FastAI/lib/python3.7/site-packages/fastai/gen_doc/nbdoc.py in doc(elt)
129 use_relative_links = False
130 elt = getattr(elt, '__func__', elt)
--> 131 md = show_doc(elt, markdown=False)
132 if is_fastai_class(elt):
133 md += f'\n\n<a href="{get_fn_link(elt)}" target="_blank" rel="noreferrer noopener">Show in docs</a>'
~/miniconda3/envs/p37cu10FastAI/lib/python3.7/site-packages/fastai/gen_doc/nbdoc.py in show_doc(elt, doc_string, full_name, arg_comments, title_level, alt_doc_string, ignore_warn, markdown, show_tests)
114 else: raise Exception(f'doc definition not supported for {full_name}')
115 source_link = get_function_source(elt) if is_fastai_class(elt) else ""
--> 116 test_link, test_modal = get_pytest_html(elt, anchor_id=anchor_id) if show_tests else ('', '')
117 title_level = ifnone(title_level, 2 if inspect.isclass(elt) else 4)
118 doc = f'<h{title_level} id="{anchor_id}" class="doc_header">{name}{source_link}{test_link}</h{title_level}>'
~/miniconda3/envs/p37cu10FastAI/lib/python3.7/site-packages/fastai/gen_doc/nbtest.py in get_pytest_html(elt, anchor_id)
55 def get_pytest_html(elt, anchor_id:str)->Tuple[str,str]:
56 md = build_tests_markdown(elt)
---> 57 html = HTMLExporter().markdown2html(md).replace('\n','') # nbconverter fails to parse markdown if it has both html and '\n'
58 anchor_id = anchor_id.replace('.', '-') + '-pytest'
59 link, body = get_pytest_card(html, anchor_id)
TypeError: markdown2html() missing 1 required positional argument: 'source'
running most recent fastAI installed into a python 3.7.3 conda environment and with jupyter notebook.
However the next cell displays the confusion matrix ok.
suggestion for resolution?
Can someone help me with understanding how to choose the value from LR Finder. From what I understood, select the max learning rate as the LR right before the reporter plot starts showing gain in loss as we keep on increasing the LR. The minimum can then be chosen as 10x or 100x smaller than max.
I understand the LR slice we chose for resnet 34 but I don’t get the slice we chose for resnet 50. Since the LR decreases until after 1e-2, shouldn’t the slice be 1e-2 to 1e-4? We have however chosen 1e-6 to 1e-4.
Fitting on the latter slice gives better results
Refer the below thread, it might help. Although it’s specific to colab, you might find workarounds for your environment -
