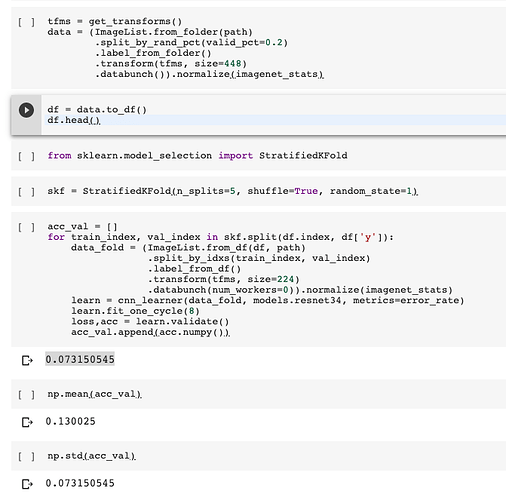
I tried to implement cross-validation in fastai, but I failed.
I have used sklearn to help generate the folds, then train a fastai classifier on each train and validation dataset combination.
Can you tell me in more detail? I divided the data set into 5 folds to train, and when I did the second iteration, I got an error.
This my way to implement Stratified K Fold cross validation with fast.ai and scikitlearn
I think that is ok and easy but I’m a newbie
have u ever raise an error : out of memory? thanks for advance!
I guess probably you can reduce the batch_size to avoid that error.
I was just asking myself: which is the purpose of cross-validation in this situation? My guess: to get a better estimate of the accuracy of the model in the test_set?
Also what about early stopping if error starts increasing in the validation set, how can we do it?
Regarding your second question you should probably have a look at the EarlyStoppingCallback in the docs.
Is the y in df[‘y’] the tags/label column in the df?
Yes, it is ![]()
Thank you muellerzr!
Hi all, how would you adapt @farconada’s method to fastai v2?
I tried playing with @muellerzr’s notebook but can’t seem to get it to work in the case of tabular data. I also only have 1 dataframe to work with (no separate validation set), so I’m finding it difficult to adapt his code. Would love to get some help from you @muellerzr!
Otherwise there is this notebook: https://github.com/muellerzr/Practical-Deep-Learning-for-Coders-2.0/blob/e5587eaacbe389df513e06407a498fd0a55897c5/Tabular%20Notebooks%20(old)/03b_kfold.ipynb
(old notebook, but only change required: from fastai.x rather than from fastai2)
Hi all,
I’m going a bit nuts with implementing KFold CV in FastAI, so I hope someone can help on this slightly older thread!
Essentially I am using FastAI V2 and am using a Tabular Learner with cont & cat cols to build a NN. But before I do that I create a robust train & unseeen test dataset:
X_train, X_test = train_test_split(all_train_data, test_size = 0.15,random_state=1)
X_train.reset_index(inplace=True,drop=True)
X_test.reset_index(inplace=True,drop=True)
Having followed: https://github.com/muellerzr/Practical-Deep-Learning-for-Coders-2.0/blob/e5587eaacbe389df513e06407a498fd0a55897c5/Tabular%20Notebooks%20(old)/03b_kfold.ipynb I then am trying to create a stratified kfold split and pass through the folds in a loop that I load train data, train the model and then aim to predict against the unseen test data (X_test) above.
However, the fun/annoyance begins when I call get_preds(dl=test_dl) and I get:
~/opt/anaconda3/envs/capstone_python/lib/python3.8/site-packages/torch/nn/functional.py in embedding(input, weight, padding_idx, max_norm, norm_type, scale_grad_by_freq, sparse)
1812 # remove once script supports set_grad_enabled
1813 _no_grad_embedding_renorm_(weight, input, max_norm, norm_type)
-> 1814 return torch.embedding(weight, input, padding_idx, scale_grad_by_freq, sparse)
1815
1816
IndexError: index out of range in self
Originally I thought that this was due to the random split nature of SKLearn’s train_test_split, however, I’ve reset the index for both train and test so can’t figure out why I’m getting this error. Is it because the learn.get_preds() can’t find an embedding element or something?
The code that runs the loop is as follows:
val_pct = L()
test_pct = L()
test_preds = L()
f1_results = []
auc_results = []
mcc_results = []
bal_acc_results = []
strat_kfold = StratifiedKFold(n_splits=5, shuffle=True)
res = strat_kfold.split(X_train.index,X_train['macro_LGA'])
tab_config = tabular_config(embed_p=0.25,use_bn=True,ps=[0.025],act_cls=Swish())
for x, y in res:
ix = (L(list(x)),L(list(y)))
train_dl = TabularDataLoaders.from_df(df=X_train,cat_names=cat_cols,
cont_names=cont_cols,y_names=dep_var,
procs=procs,bs=64,splits=ix)
tab_config = tabular_config(embed_p=0.25,use_bn=True,ps=[0.025,0.015],act_cls=Mish())
learn = tabular_learner(train_dl,config=tab_config,loss_func=loss_func,
metrics=[accuracy,roc_auc],lr=0.00005,emb_szs=emb_szs, layers=[100,50],
opt_func=RAdam)
learn.fit(n_epoch=3, cbs=SaveModelCallback(monitor='valid_loss'))
test_dl = TabularPandas(X_test, procs=procs, cat_names=cat_cols,
cont_names=cont_cols, y_names="macro_LGA")
test_dl = TabDataLoader(test_dl)
test_pct.append(learn.get_preds(dl=test_dl)[0])
Help?!
Thanks!