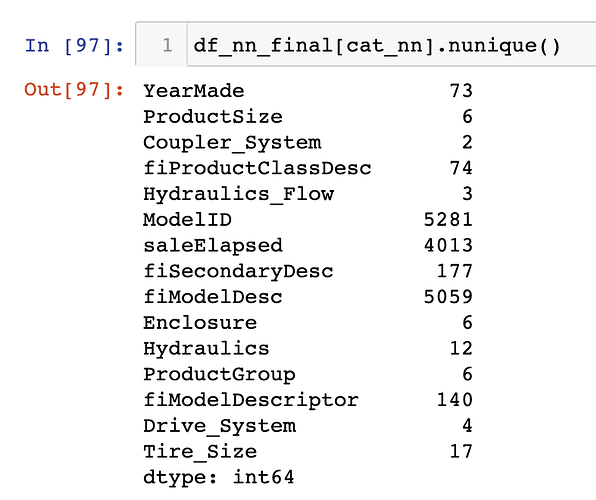
Towards the end of the chapter, we see the following paragraph:
The fact that there are two variables pertaining to the “model” of the equipment, both with similar very high cardinalities, suggests that they may contain similar, redundant information… Having a column with 5,000 levels means needing 5,000 columns in our embedding matrix, which would be nice to avoid if possible. Let’s see what the impact of removing one of these model columns has on the random forest:
Then, we
xs_filt2 = xs_filt.drop('fiModelDescriptor', axis=1)
valid_xs_time2 = valid_xs_time.drop('fiModelDescriptor', axis=1)
m2 = rf(xs_filt2, y_filt)
m_rmse(m2, xs_filt2, y_filt), m_rmse(m2, valid_xs_time2, valid_y)
=> (0.176853, 0.229973)
cat_nn.remove('fiModelDescriptor')
My question is, if the columns which create the need for larger and more memory-intensive embedding matrices are the columns with cardinalities around 5,000+ (i.e. ModelID and fiModelDesc ), why is it that we instead remove a column which has a cardinality of only 140, i.e. fiModelDescriptor? Why not remove one of the first two?