Hi @jeremy,
At first, I would like to thank you for the great lecture yesterday. After the session, I was looking at the fastai docs https://docs.fast.ai/index.html and found it very useful in the context of understanding how fastai works. Since I am now actively working on Image Regression problems now, I was checking the document feature for loading and training Image Regression but I couldn’t find any about it. Is there any thread or sample snippet where I can train image regression problems in fastai. I am a stalwart of the concepts of one-cycle learning rate techniques. I would love to train my model using fastai.
Can you link or give an example of image regression? I haven’t heard that term before, but maybe with an example I would be able to help point you in the right direction.
There are multiple scenarios. Given an image say of a person estimate height, age, weight etc. density estimation would be another.
Thanks for the response @KevinB. An apt example of image regression would be of a below scenario. Assume that images are labelled with a particular numerical score(eg. explicit user ratings). Then you can build a deep learning network to predict the quality rating given the image. In the previous libraries, there is an attribute called is_reg in the ConvnetBuilder() function which will actually change the final layers of the Convolutional Model from softmax to Linear. Kindly check the below snippet.
Also you can find this code here https://github.com/fastai/fastai/blob/master/old/fastai/conv_learner.py
This particular feature is actually missing in the version 3
Am looking at something very similar. Have done it in Keras. just started exploring fastai. As a first step, I one guess would have to modify the databunch so that generated labels are ints/floats rather than OHE. And the custom head would be a single neuron with sigmoid activation whose output is scaled to the range you are trying to predict. And finally use MSE/MAE loss function.
You might be able to use it out of the box.
learn = ConvLearner(data, models.resnet34, metrics=error_rate)
gives a linear final layer and doesn’t have a softmax layer at the end. Then when you want to actually get predictions from the learner, I wonder if you could do something on the interpreter side (Not 100% sure if that’s how it’s designed)
interp = ClassificationInterpretation.from_learner(learn, sigmoid=False)
This would give you that linear layer of predictions out from the initial ConvLearner. Is this helpful at all? FYI here is what the ConvLearner from above looks like when I do learn.model
ConvLearner(data=<fastai.vision.data.ImageDataBunch object at 0x7f74aa5dd710>, model=Sequential(
(0): Sequential(
(0): Conv2d(3, 64, kernel_size=(7, 7), stride=(2, 2), padding=(3, 3), bias=False)
(1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace)
(3): MaxPool2d(kernel_size=3, stride=2, padding=1, dilation=1, ceil_mode=False)
(4): Sequential(
(0): BasicBlock(
(conv1): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(1): BasicBlock(
(conv1): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(2): BasicBlock(
(conv1): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
)
(5): Sequential(
(0): BasicBlock(
(conv1): Conv2d(64, 128, kernel_size=(3, 3), stride=(2, 2), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(downsample): Sequential(
(0): Conv2d(64, 128, kernel_size=(1, 1), stride=(2, 2), bias=False)
(1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
)
(1): BasicBlock(
(conv1): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(2): BasicBlock(
(conv1): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(3): BasicBlock(
(conv1): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
)
(6): Sequential(
(0): BasicBlock(
(conv1): Conv2d(128, 256, kernel_size=(3, 3), stride=(2, 2), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(downsample): Sequential(
(0): Conv2d(128, 256, kernel_size=(1, 1), stride=(2, 2), bias=False)
(1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
)
(1): BasicBlock(
(conv1): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(2): BasicBlock(
(conv1): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(3): BasicBlock(
(conv1): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(4): BasicBlock(
(conv1): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(5): BasicBlock(
(conv1): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
)
(7): Sequential(
(0): BasicBlock(
(conv1): Conv2d(256, 512, kernel_size=(3, 3), stride=(2, 2), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(downsample): Sequential(
(0): Conv2d(256, 512, kernel_size=(1, 1), stride=(2, 2), bias=False)
(1): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
)
(1): BasicBlock(
(conv1): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
(2): BasicBlock(
(conv1): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn1): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(relu): ReLU(inplace)
(conv2): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(bn2): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
)
)
)
(1): Sequential(
(0): AdaptiveConcatPool2d(
(ap): AdaptiveAvgPool2d(output_size=1)
(mp): AdaptiveMaxPool2d(output_size=1)
)
(1): Lambda()
(2): BatchNorm1d(1024, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(3): Dropout(p=0.25)
(4): Linear(in_features=1024, out_features=512, bias=True)
(5): ReLU(inplace)
(6): BatchNorm1d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(7): Dropout(p=0.5)
(8): Linear(in_features=512, out_features=37, bias=True)
)
), opt_func=functools.partial(<class 'torch.optim.adam.Adam'>, betas=(0.9, 0.99)), loss_func=<function cross_entropy at 0x7f74b2249ae8>, metrics=[<function error_rate at 0x7f74ab804730>], true_wd=True, bn_wd=True, wd=0.01, train_bn=True, path=PosixPath('/home/kbird/.fastai/data/oxford-iiit-pet/images'), model_dir='models', callback_fns=[<class 'fastai.basic_train.Recorder'>], callbacks=[], layer_groups=[Sequential(
(0): Conv2d(3, 64, kernel_size=(7, 7), stride=(2, 2), padding=(3, 3), bias=False)
(1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace)
(3): MaxPool2d(kernel_size=3, stride=2, padding=1, dilation=1, ceil_mode=False)
(4): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(5): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(6): ReLU(inplace)
(7): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(8): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(9): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(10): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(11): ReLU(inplace)
(12): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(13): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(14): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(15): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(16): ReLU(inplace)
(17): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(18): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(19): Conv2d(64, 128, kernel_size=(3, 3), stride=(2, 2), padding=(1, 1), bias=False)
(20): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(21): ReLU(inplace)
(22): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(23): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(24): Conv2d(64, 128, kernel_size=(1, 1), stride=(2, 2), bias=False)
(25): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(26): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(27): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(28): ReLU(inplace)
(29): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(30): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(31): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(32): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(33): ReLU(inplace)
(34): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(35): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(36): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(37): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(38): ReLU(inplace)
(39): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(40): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
), Sequential(
(0): Conv2d(128, 256, kernel_size=(3, 3), stride=(2, 2), padding=(1, 1), bias=False)
(1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace)
(3): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(4): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): Conv2d(128, 256, kernel_size=(1, 1), stride=(2, 2), bias=False)
(6): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(7): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(8): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(9): ReLU(inplace)
(10): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(11): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(12): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(13): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(14): ReLU(inplace)
(15): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(16): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(17): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(18): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(19): ReLU(inplace)
(20): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(21): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(22): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(23): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(24): ReLU(inplace)
(25): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(26): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(27): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(28): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(29): ReLU(inplace)
(30): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(31): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(32): Conv2d(256, 512, kernel_size=(3, 3), stride=(2, 2), padding=(1, 1), bias=False)
(33): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(34): ReLU(inplace)
(35): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(36): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(37): Conv2d(256, 512, kernel_size=(1, 1), stride=(2, 2), bias=False)
(38): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(39): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(40): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(41): ReLU(inplace)
(42): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(43): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(44): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(45): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(46): ReLU(inplace)
(47): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1), bias=False)
(48): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
), Sequential(
(0): AdaptiveAvgPool2d(output_size=1)
(1): AdaptiveMaxPool2d(output_size=1)
(2): Lambda()
(3): BatchNorm1d(1024, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(4): Dropout(p=0.25)
(5): Linear(in_features=1024, out_features=512, bias=True)
(6): ReLU(inplace)
(7): BatchNorm1d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(8): Dropout(p=0.5)
(9): Linear(in_features=512, out_features=37, bias=True)
)])
My guess is you probably wouldn’t want that last 512 -> 37 linear layer probably?
Thanks arul. This is interesting. No activation. I had read somewhere the better way to to go would be to constrain the output between 0-1 using sigmoid. And if your predicted range was between some min and max, then multiply the sigmoid output with (max-min) and then add min.
Hi kevin,
I am not sure it would work out of the box. t
You are right. The out_features would have to be 1.
But both metrics and data would have to be modified. I only see RMSPE metric related to regression. may have to write custom loss function. and also modify databunch.
Thanks for the explanation @KevinB. This might be useful for me to go ahead with the training. Yes I might have to remove the final output features to 1 rather than 37 (I might have to override or explicitly change the final layers and the loss function to MSELoss() or L1Loss() . All these features were handled very well in the previous version thought ![]()
Yeah, I am currently looking into the loss function from lesson 1 and I am really confused at where the loss function is actually defined. (I believe it is coming from the databunch. I’ll let you know if I find a good way to modify that loss function.
The loss function has been handled in the previous version as below. It would be helpful if we have this feature in the new version.
Great to know that even you are looking out for the same problem. We can connect and discuss if you are interested in solving this problem forward.
Would love to. Will have to bear with me though. Trying to get a hang of the fastai/pytorch framework. Quite a bit different from Keras. For example activation function being tied to loss feels so different to me. Trying to come upt speed
I had a similar problem and wrote a quick guide on how I resolved it: https://medium.com/@btahir/a-quick-guide-to-using-regression-with-image-data-in-fastai-117304c0af90
Hopefully it helps!
Hi, would anyone happen to know if progress has been made in implementing image regression? I have learned the trick for setting label_class=FloatList in the data loader, however, my problem is that the model wants to predict outside of the bounds of my training data. Constraining a sigmoid between a min and max value is the approach Jeremy advocates in the collaborative filtering model. Can anyone point me to an example of this for a cnn_learner?
Here is a place in the documentation that shows reading a dataset with the goal of image regression: https://docs.fast.ai/tutorial.data.html#A-regression-example
Thanks, yes I’ve seen that particular example. It doesn’t provide a mechanism to constrain the range of predictions, however. The tabular learners have a y_range argument, but the cnn’s do not. My problem is that my image regression models predict outside of the range of values in my training data.
What I have done is simply to put a sigmoid in front of the MSELoss. It smoothly squishes the activations into [0,1], and the model trains using this constraint. If your training labels have a different range, multiply and add to sigmoid in order to map to that range.
def myMSE(input,target):
return F.mse_loss(torch.sigmoid(input), target)
learn.loss_func = myMSE
Then when you get the prediction you’ll also apply sigmoid to convert activations to predictions.
I suppose you could equivalently append a sigmoid activation layer to the head of the model.
HTH,
Malcolm
Thanks for your response, I have tried the approach of modifying the loss function you propose. My data are indeed scaled 0 to 1. I’m getting better results with the stock loss function.
Perhaps I am not implementing correctly when you say: “apply sigmoid to convert activations to predictions”
Presuming I am applying TTA at the end, is this what you are suggesting:
yhat, y = learn.TTA()
torch.sigmoid(yhat)
Would I expect different results from a custom head approach?
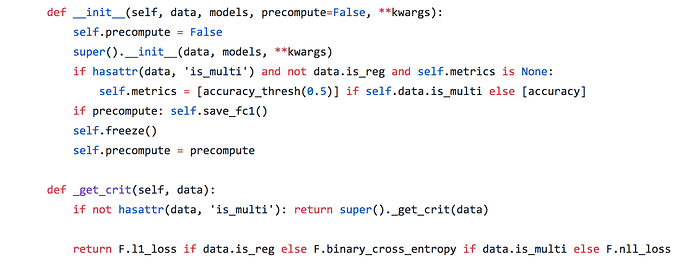
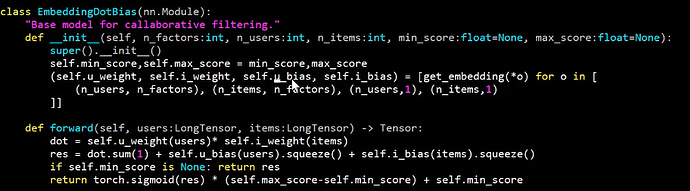
Referring to lesson 4 in which Jeremy discusses collaborative filtering to predict movie rating between 0 and 5, the sigmoid applied at the end is defined:
Where I suspect the relevant line for scaling the data to max and min of y is:
torch.sigmoid(res) * (self.max_score-self.min_score) + self.min_score
If I wanted to pursue a custom head approach, how could I include the above? I’m not finding a whole lot of info on custom head. When I print out the model with the stock head, I see:
Will all the elements 0:9 need to be replaced or only element 9?
Good morning.
Presuming I am applying TTA at the end, is this what you are suggesting:
yhat, y = learn.TTA()
torch.sigmoid(yhat)
No, I meant learn.get_preds and then apply sigmoid to convert activations to probabilities. Watch out - sometimes get_preds returns activations and other times probabilities.(Depending I think on whether it recognizes your loss function.) You can either look at the results (probabilities are never negative) or trace the code.
I am not sure what learn.TTA returns in this case, but you could trace it and find out.
Would I expect different results from a custom head approach?
I don’t know, try it! First make your own expectation, try the experiment, and use the actual outcome to update and refine your understanding.
If you’ll allow me to give some advice, speaking only as an “advanced beginner”: install PyTorch or VSCode and start tracing fastai and PyTorch to see what actually happens. The code is quite advanced in places, but you can always step over a line and see its effect on variables. Forget PDB for now - to explore well you will need a competent debugger with stepping, variable and code context, and code navigation.
Taking on a Kaggle competition was the best choice I ever made for this course, because it forced me to start actually experimenting and tracing code. My own learning rate about machine learning and fastai shot up 10x. It’s certainly not easy to jump into areas that look totally obscure, but with engaged effort bit by bit things become clear. Asking others’ advice and opinions can only take you so far - you have to actually wrestle with the material.
Where I suspect the relevant line for scaling the data to max and min of y is:
torch.sigmoid(res) * (self.max_score-self.min_score) + self.min_score
Right - this is the line that scales the model’s output. You might copy it into a notebook and experiment with parameters until you are fully confident about how it works.
If I wanted to pursue a custom head approach, how could I include the above? I’m not finding a whole lot of info on custom head. When I print out the model with the stock head, I see:…
Will all the elements 0:9 need to be replaced or only element 9?
To better understand custom heads you should trace through the code for cnn_learner to see how the head is made and attached to the backbone. Even if you don’t fully understand the library code you can still copy fragments into a notebook to experiment with and use by analogy.
Here you would append an nn.Sigmoid layer after the last Linear layer. The cnn_learner library code demonstrates how to construct a sequence of layers. To use a scaled sigmoid, google for examples of custom nn.Module, then write your own by analogy. The scaling would be done in the forward() method. Then instead of nn.Sigmoid, append your custom Module.
Good luck! Soon enough you will be joining me in higher level confusions. ![]()