New user here, adding to this conversation as I couldn’t see how to start a new thread…
I think I’ve got a similar issue to the one above - essentially I have a Tabular model that’s training well with an RMSE of ~2.0.
Before submitting to a competition, I’m using re-adding the entire training and validation set as a test set, then using get_preds against this (to ensure I’ve got the workflow functioning properly). However when I export that model, load it back in and get_preds against that test set, the RMSE is around 60!
Would really appreciate if someone could help me out, and point out where I’m going wrong - this is a massive difference. I’ve included my code and some of the differences in results below:
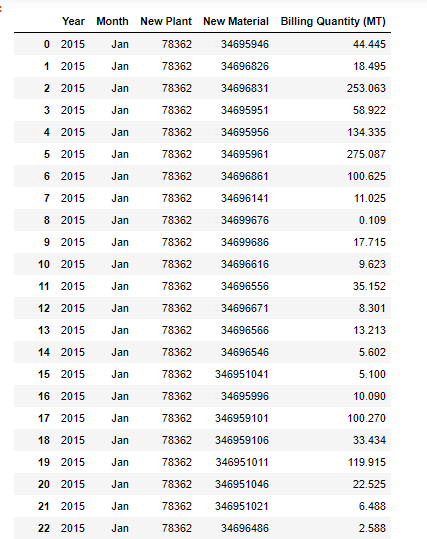
Drop timestamps, rename target and declare continuous vs categorical variables for fast.ai
cleaned_df = merged_df.drop(columns=[‘target_timestamp’]).rename(columns={
‘target_PPO:AC4_1A:TIC7201-PV’: ‘target’
})
for i in range(0, len(cleaned_df.columns)):
cleaned_df.iloc[:,i] = pd.to_numeric(cleaned_df.iloc[:,i], errors=‘ignore’)
cont_vars = cleaned_df.columns.values.tolist()
cont_vars.remove(‘timestamp’)
add_datepart(cleaned_df, “timestamp”, drop=True)
cat_vars = []
valid_idx = range(int(.7len(cleaned_df)), int(.9len(cleaned_df)))
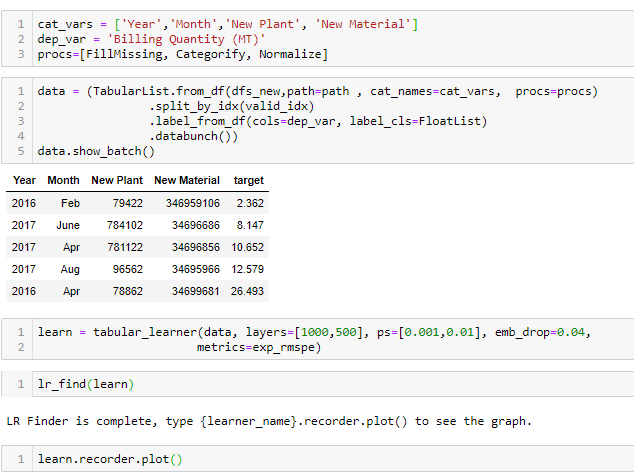
procs=[FillMissing, Categorify, Normalize]
dep_var = ‘target’
Create indexed databunch
data = (TabularList.from_df(cleaned_df, path=data_path, cat_names=cat_vars, cont_names=cont_vars, procs=procs)
.split_by_idx(valid_idx)
.label_from_df(cols=dep_var, label_cls=FloatList, log=False)
.databunch())
max_y = (np.max(cleaned_df[‘target’])*1.2)
y_range = torch.tensor([0, max_y], device=defaults.device)
learn = tabular_learner(data, layers=[1000,500], ps=[0.01,0.1], emb_drop=0.04,
y_range=y_range, metrics=root_mean_squared_error)
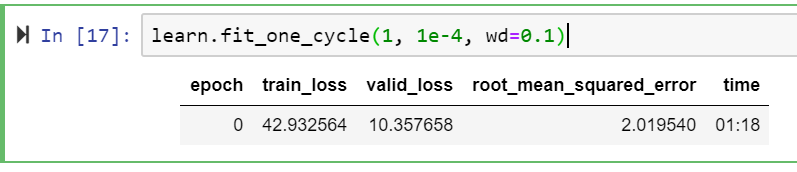
learn.fit_one_cycle(1, 1e-4, wd=0.1)
test_df = merged_df.drop(columns=[‘timestamp’, ‘target_timestamp’,‘target_PPO:AC4_1A:TIC7201-PV’])
test_df[‘target’] = np.arange(len(test_df))
for i in range(0, len(test_df.columns)):
test_df.iloc[:,i] = pd.to_numeric(test_df.iloc[:,i], errors=‘ignore’)
model_name = ‘regressor’
test_learn = load_learner(path=data_path, file=f’{model_name}.pkl’,
test=TabularList.from_df(test_df, cat_names=cat_vars, cont_names=cont_vars))
preds, y = test_learn.get_preds(DatasetType.Test)
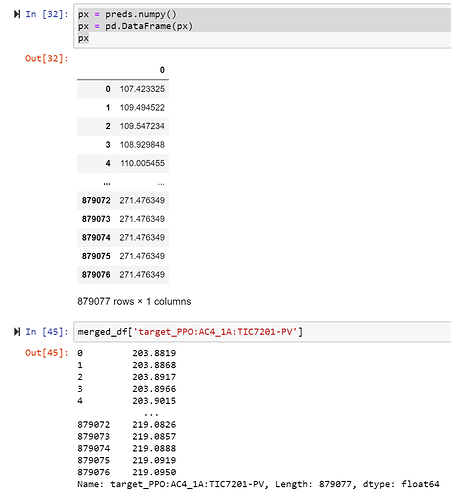
px = preds.numpy()
px = pd.DataFrame(px)
px
testing = pd.merge(merged_df, px, left_index=True, right_index=True)
testing[‘target_PPO:AC4_1A:TIC7201-PV’] = testing[‘target_PPO:AC4_1A:TIC7201-PV’].astype(‘float’)
Calculate RMSE
((testing[‘target_PPO:AC4_1A:TIC7201-PV’] - testing.iloc[73]) ** 2).mean() ** .5
Result: 59.336321997070314