Wiki topic to discuss fastai v2 tabular module.
NOTE: Below is not the best way to read code, it’s rather checking the internals. I will keep the post as is, as this is a rabbit hole that hopefully you won’t fall into 
Let’s set it up with Setup
Alright then, after code walk-thru 8, its time to explore Tabular!
So after some exploration, the first thing that I feel we need to look at is setup. After all, setup helps us set up for training. 
There’s also a little bit of a need to understand __mro__ and Super calls which will, in general, help us understand the codebase better. Recently, I’ve been spending a lot of time reading official docs and various articles, here are a couple of recommendations:
Alright then, hopefully, you’ve read the above articles, in particular, the one that explains Super and __mro__.
So let’s get started with 40_tabular_core.ipynb.
According to what I’ve understood, there are two main things inside Tabular:
- The Tabular Object
toitself - The Tabular Processes ie.,
TabularProcsuch asCategorifyetc
Tabular Object TabularPandas
First things first - let’s check the __mro__.
TabularPandas.__mro__
>>>
(__main__.TabularPandas,
__main__.Tabular,
local.core.CollBase,
local.core.GetAttr,
local.core.BaseObj,
object)
Next step, let’s see where setup is, because, this is where we first call setup like so:
df = pd.DataFrame({'a':[0,1,2,0,2]})
to = TabularPandas(df, Categorify, 'a')
to.setup()
As you can see, we call setup on Tabular Object to therefore, it’s important to understand where it get’s called from.
Since, setup is not directly defined inside TabularPandas we go up the __mro__ and check inside Tabular where we do find setup defined as:
def setup(self):
self.procs.setup(self)
Ahan! So this in turn will call procs.setup and remember from walk thru-8 procs is a Pipeline of Transforms that we pass such as Categorify!
So, this will call the Pipeline’s setup which in turn calls the setup on each of the transforms.
def setup(self, items=None):
print("I believe I was called")
self.items = items
tfms,self.fs = self.fs,L()
for t in tfms: self.add(t,items)
def add(self,t, items=None):
t.setup(items)
self.fs.append(t)
So let’s check where setup is defined inside Categorify.
Categorify has setups but not setup so this can’t be it! Let’s check its __mro__.
Categorify.__mro__
>>>
(__main__.Categorify,
__main__.TabularProc,
local.data.transform.InplaceTransform,
local.data.transform.Transform,
object)
So next place to check is TabularProc which has setup which first calls super().setup(items) and then returns based on that. Okay, so what is super then? Let’s check in __mro__. It can’t be InplaceTranform because there is no super inside it. So it has to be Transform which is defined as:
def setup(self, items=None): return self.setups(items)
So it in turn calls setups which should be Categorify setups.
And that is how we end up with a CateogryMap like so
def setups(self, dsrc):
self.classes = {n:CategoryMap(getattr(dsrc,'train',dsrc).iloc[:,n].items, add_na=True) for n in dsrc.all_cat_names}
** Writing such posts is a great idea - it helps me deepen my understanding and hopefully helps others along the way! You should do it too! 
** As usual, please feel free to correct me, since I am very new to this myself. Thanks for the feedback!
** I am also a little overwhelmed by the number of layers and base classes, steps we had to go through to call setup.
@arora_aman that’s a rather complicated way of reading the code - it would be like understanding how to use Python by debugging its internals!
Instead, I’d suggest learning the key bits of the API, just like you learn the functionality of Python based on what it does, rather than how it’s implemented.
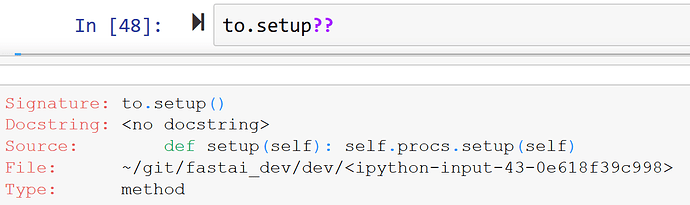
In Python rather than looking at the mro, instead it’s generally easier just to look to see what a class inherits from. Or, easier still, just ask Jupyter what’s being called:
A Pipeline is a list of Transforms which are applied through function composition. A Transform can define 3 things: __call__, decode, and setup, which call encodes, decodes, and setups respectively. They are called using TypeDispatch and applied to each item in a tuple (if as_item is true).
I’m attempting to make a starter kernel for the Ashrae Energy Prediction Kaggle Competition, and it seems I’m running into a few errors. I’m using the standard tabular library instead of rapids as I’m having issues conda installing in a kernel. I have cat, cont vars, procs, and splits defined as:
cat_vars = ['primary_use']
cont_vars = ['square_feet', 'year_built']
procs = [Normalize, Categorify, FillMissing]
splits = RandomSplitter()(range_of(train))
When I try to create our TabularPandas object as so:
to = TabularPandas(train, procs, cat_vars, cont_vars, y_names="meter_reading", splits=splits)
I get an error pointing back to the transform.py:
TypeError Traceback (most recent call last)
<ipython-input-37-d9389de6d074> in <module>
----> 1 to = TabularPandas(train, procs, cat_vars, cont_vars, y_names="meter_reading", splits=splits)
/kaggle/working/fastai_dev/dev/local/tabular/core.py in __init__(self, df, procs, cat_names, cont_names, y_names, is_y_cat, splits, do_setup)
35
36 store_attr(self, 'y_names,is_y_cat')
---> 37 self.cat_names,self.cont_names,self.procs = L(cat_names),L(cont_names),Pipeline(procs, as_item=True)
38 self.cat_y = None if not is_y_cat else y_names
39 self.cont_y = None if is_y_cat else y_names
/kaggle/working/fastai_dev/dev/local/core/transform.py in __init__(self, funcs, as_item, split_idx)
177 else:
178 if isinstance(funcs, Transform): funcs = [funcs]
--> 179 self.fs = L(ifnone(funcs,[noop])).map(mk_transform).sorted(key='order')
180 for f in self.fs:
181 name = camel2snake(type(f).__name__)
/kaggle/working/fastai_dev/dev/local/core/foundation.py in map(self, f, *args, **kwargs)
338 else f.format if isinstance(f,str)
339 else f.__getitem__)
--> 340 return self._new(map(g, self))
341
342 def filter(self, f, negate=False, **kwargs):
/kaggle/working/fastai_dev/dev/local/core/foundation.py in _new(self, items, *args, **kwargs)
292 super().__init__(items)
293
--> 294 def _new(self, items, *args, **kwargs): return type(self)(items, *args, use_list=None, **kwargs)
295 def __getitem__(self, idx): return self._get(idx) if is_indexer(idx) else L(self._get(idx), use_list=None)
296
/kaggle/working/fastai_dev/dev/local/core/foundation.py in __call__(cls, x, *args, **kwargs)
40 return x
41
---> 42 res = super().__call__(*((x,) + args), **kwargs)
43 res._newchk = 0
44 return res
/kaggle/working/fastai_dev/dev/local/core/foundation.py in __init__(self, items, use_list, match, *rest)
285 if items is None: items = []
286 if (use_list is not None) or not _is_array(items):
--> 287 items = list(items) if use_list else _listify(items)
288 if match is not None:
289 if is_coll(match): match = len(match)
/kaggle/working/fastai_dev/dev/local/core/foundation.py in _listify(o)
221 if isinstance(o, list): return o
222 if isinstance(o, str) or _is_array(o): return [o]
--> 223 if is_iter(o): return list(o)
224 return [o]
225
/kaggle/working/fastai_dev/dev/local/core/foundation.py in __call__(self, *args, **kwargs)
193 if isinstance(v,_Arg): kwargs[k] = args.pop(v.i)
194 fargs = [args[x.i] if isinstance(x, _Arg) else x for x in self.pargs] + args[self.maxi+1:]
--> 195 return self.fn(*fargs, **kwargs)
196
197 #Cell
/kaggle/working/fastai_dev/dev/local/core/transform.py in mk_transform(f, as_item)
152 def mk_transform(f, as_item=True):
153 "Convert function `f` to `Transform` if it isn't already one"
--> 154 f = instantiate(f)
155 return f if isinstance(f,Transform) else Transform(f, as_item=as_item)
156
/kaggle/working/fastai_dev/dev/local/core/utils.py in instantiate(t)
389 def instantiate(t):
390 "Instantiate `t` if it's a type, otherwise do nothing"
--> 391 return t() if isinstance(t, type) else t
392
393 #Cell
/kaggle/working/fastai_dev/dev/local/core/transform.py in __call__(cls, *args, **kwargs)
60 getattr(cls,n).add(f)
61 return f
---> 62 return super().__call__(*args, **kwargs)
63
64 @classmethod
TypeError: __init__() missing 2 required positional arguments: 'mean' and 'std'
Stating that for Normalize (I’m assuming normalize since it’s mean and std) it could not grab the mean and standard deviation. How should I go about debugging/fixing this? 
There is a conflict of names here. You probably have the Normalize from vision interfering with the Normalize from tabular.
Interesting. I’ll try a fresh restart and importing again. Will let you know if that fixed the issue. Thanks! 
That seems to have done the trick! I’ll post a starter Kernel for Kaggle here shortly 
Here is my notebook getting this working for the Kaggle Competition  It was infact an import issue.
It was infact an import issue. The only problem I’m noticing is kaggle does not want to export/commit my work due to the but let me know if anyone has questions or ideas for improvement! (I made it quickly as a tutorial, and I’m unsure as to why val_loss is infinite) here is the kernel if you prefer Kagglegit clone so I’m unsure what to do
Would it be feasible to persist o2i? I have a use case where I would add new data on a daily basis, with additional categorical features. If the creation of o2i is changed to updating a pickled o2i, this would be friendlier for production.
I tried building a cat to int dict and using it to merge as part of the pre-processing, with a ‘if is_numerical_dtype’ line added to Category_Map in 06_data_transforms, but sticking with o2i seems less hacky.
@sgugger I’m attempting to try out a test set using the adults dataset. Current setup:
to = TabularPandas(df_main, procs, cat_names, cont_names, y_names="salary", splits=splits)
to_test = TabularPandas(df_test, procs, cat_names, cont_names, y_names="salary", splits=None)
dbch = to.databunch()
tst = test_dl(dbch, to_test)
To generate the test dataloader. Then I do
learn.get_preds(dl=tst)
I get an AttributeError:
---------------------------------------------------------------------------
AttributeError Traceback (most recent call last)
<ipython-input-15-524ee7bdf73c> in <module>()
----> 1 learn.get_preds(dl=tst)
10 frames
/usr/local/lib/python3.6/dist-packages/fastai2/learner.py in get_preds(self, ds_idx, dl, with_input, with_loss, with_decoded, act)
257 with self.no_logging(), self.added_cbs(cb), self.loss_not_reduced():
258 self(_before_epoch)
--> 259 self._do_epoch_validate(ds_idx, dl)
260 self(_after_epoch)
261 if act is None: act = getattr(self.loss_func, 'activation', noop)
/usr/local/lib/python3.6/dist-packages/fastai2/learner.py in _do_epoch_validate(self, ds_idx, dl)
221 try:
222 self.dl = dl; self('begin_validate')
--> 223 with torch.no_grad(): self.all_batches()
224 except CancelValidException: self('after_cancel_validate')
225 finally: self('after_validate')
/usr/local/lib/python3.6/dist-packages/fastai2/learner.py in all_batches(self)
191 def all_batches(self):
192 self.n_iter = len(self.dl)
--> 193 for o in enumerate(self.dl): self.one_batch(*o)
194
195 def one_batch(self, i, b):
/usr/local/lib/python3.6/dist-packages/fastai2/data/load.py in __iter__(self)
90 self.randomize()
91 self.before_iter()
---> 92 for b in _loaders[self.fake_l.num_workers==0](self.fake_l): yield self.after_batch(b)
93 self.after_iter()
94
/usr/local/lib/python3.6/dist-packages/fastai2/core/transform.py in __call__(self, o)
198 self.fs.append(t)
199
--> 200 def __call__(self, o): return compose_tfms(o, tfms=self.fs, split_idx=self.split_idx)
201 def __repr__(self): return f"Pipeline: {self.fs}"
202 def __getitem__(self,i): return self.fs[i]
/usr/local/lib/python3.6/dist-packages/fastai2/core/transform.py in compose_tfms(x, tfms, is_enc, reverse, **kwargs)
146 for f in tfms:
147 if not is_enc: f = f.decode
--> 148 x = f(x, **kwargs)
149 return x
150
/usr/local/lib/python3.6/dist-packages/fastai2/core/transform.py in __call__(self, x, **kwargs)
84 @property
85 def use_as_item(self): return ifnone(self.as_item_force, self.as_item)
---> 86 def __call__(self, x, **kwargs): return self._call('encodes', x, **kwargs)
87 def decode (self, x, **kwargs): return self._call('decodes', x, **kwargs)
88 def setup(self, items=None): return self.setups(items)
/usr/local/lib/python3.6/dist-packages/fastai2/core/transform.py in _call(self, fn, x, split_idx, **kwargs)
92 if split_idx!=self.split_idx and self.split_idx is not None: return x
93 f = getattr(self, fn)
---> 94 if self.use_as_item or not is_listy(x): return self._do_call(f, x, **kwargs)
95 res = tuple(self._do_call(f, x_, **kwargs) for x_ in x)
96 return retain_type(res, x)
/usr/local/lib/python3.6/dist-packages/fastai2/core/transform.py in _do_call(self, f, x, **kwargs)
97
98 def _do_call(self, f, x, **kwargs):
---> 99 return x if f is None else retain_type(f(x, **kwargs), x, f.returns_none(x))
100
101 add_docs(Transform, decode="Delegate to `decodes` to undo transform", setup="Delegate to `setups` to set up transform")
/usr/local/lib/python3.6/dist-packages/fastai2/core/dispatch.py in __call__(self, *args, **kwargs)
96 if not f: return args[0]
97 if self.inst is not None: f = types.MethodType(f, self.inst)
---> 98 return f(*args, **kwargs)
99
100 def __get__(self, inst, owner):
/usr/local/lib/python3.6/dist-packages/fastai2/tabular/core.py in encodes(self, to)
136 def __init__(self, to): self.to = to
137 # TODO: use float for cont targ
--> 138 def encodes(self, to): return tensor(to.cats).long(),tensor(to.conts).float(), tensor(to.targ).long()
139
140 def decodes(self, o):
AttributeError: 'tuple' object has no attribute 'cats'
Any suggestions?
I can do tst.cats just fine
Perhaps the first transform is returning a Tuplr but it needs to return a tabular object which has cats and conts?
Just to make sure I tried res = ReadTabBatch(tst).encodes(tst) and it successfully returned everything.
test_dl is expecting a list of items, not a TabularPandas object. From your to_test, just build a databunch with
dbunch_test = to_test.databunch
then you can use dbunch_test.train_dl. Or dbunch_test.train_dl.new(shuffle=False, drop_last=False) for a non-shuffled and complete version (since by default shuffle and drop_last are True on the training dataloader).
I tried this with:
dbunch_test = to_test.databunch(shuffle_train=False)
tst = test_dl(dbch, dbunch_test.train_dl)
learn.get_preds(dl=tst)
(and also exactly the method you described above too)
and I still get the "AttributeError ‘tuple’ object has no attribute ‘cats’
No, you should directly use the dataloader you get, not the test_dl function.
Ah, I understand now. Thank you!!!
SUPER excited about how we can pass in labelled test sets in now and run learn.validate() on them. Thank you for this!
Yes, I know it was something that a lot of people wanted in v1 
While looking at it I realized a much simpler step:
to_test = TabularPandas(df_test, procs, cat_names, cont_names, y_names="salary")
test_dl = TabDataLoader(to_test, bs=128, shuffle=False, drop_last=False)
learn.validate(dl=test_dl)
I was wondering if it wouldn’t be better to use dbunch_test.train_dl all the way. Therefore, we avoid instanciating and using test_dl:
test_dl = TabDataLoader(to_test, bs=128, shuffle=False, drop_last=False)
In that case, we will have both
learn.validate(dl=dbunch_test.train_dl) and
preds = learn.get_preds(dl=dbunch_test.train_dl) instead of
learn.validate(dl=test_dl) and
preds = learn.get_preds(dl=dbunch_test.train_dl)
Invoking learn.validate(dl=dbunch_test.train_dl) returns the same result as learn.validate(dl=test_dl)
Also, dbunch_test.train_dl won’t drop any samples because it has drop_last=False as test_dl
The reason is because in the (FilteredBase) databunch() method we have:
dls = [dl_type(self.subset(i), bs=b, shuffle=s, drop_last=s, n=n if i==0 else None, **kwargs, **dk)
Where the s value is equal to shuffle_train which is set to False in this case.
Am I missing something regarding the need to use test_dl in this case?
I decided the second method I posted just above because it’s a lot less clunkier, and instead of generating a full DataBunch we just generate a DataLoader. (as that’s what it wants!)
Because originally (including the drop_last) it would be this:
to_test = TabularPandas(df_test, procs, cat_names, cont_names, y_names="salary")
dbunch_test = to_test.databuch()
dbunch_test.train_dl = dbunch_test.train_dl.new(shuffle=False, drop_last=False)
learn.validate(dbunch_test.train_dl)
Verses:
to_test = TabularPandas(df_test, procs, cat_names, cont_names, y_names="salary")
test_dl = TabDataLoader(to_test, bs=128, shuffle=False, drop_last=False)
learn.validate(dl=test_dl)
The second is much easier to read IMO and easier to not mess up (accidentally forgetting shuffle=False, etc). Thoughts?
Also do note the test_dl function is not what you want to use as sgugger mentioned above. Just use a regular DataLoader
My comment was prompted after I saw that you are using both test_dl and dbch_test in your Test Sets in v2.ipynb notebook.
test_dl = TabDataLoader(to_test, bs=128, shuffle=False, drop_last=False) and
dbch_test = to_test.databunch(shuffle_train=False)
I agree with you, if you don’t need the dbch_test then using test_dl alone is more explicit.
This
is not necessary anymore since we will get the same result just by writing:
dbch_test = to_test.databunch(shuffle_train=False)
dbunch_test.train_dl will autommatically have (shuffle=False, drop_last=False)