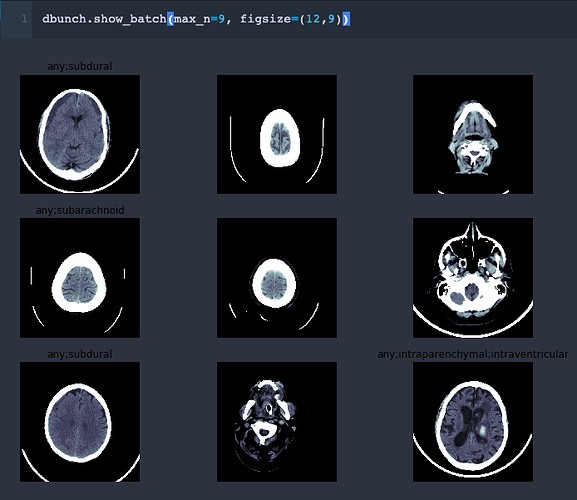
I think I’ve got it. It’s actually pretty simple.
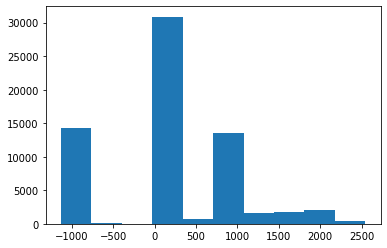
Once we flatten and sort the image pixels, we get a 1-D torch tensor which has values like tensor([-2157., -2157., -2155., ..., 1464., 1520., 1521.])
Next, we just make selection points or positions from this array and store in t like so:
imsd = self.view(-1).sort()[0]
t = torch.cat([tensor([0.001]),
torch.arange(n_bins).float()/n_bins+(1/2/n_bins),
tensor([0.999])])
t = (len(imsd)*t).long()
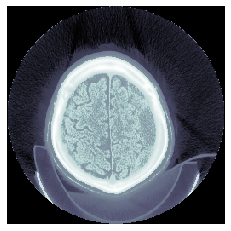
Finally we just select the pixel values from these t positions from the image.
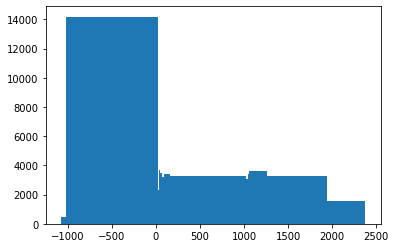
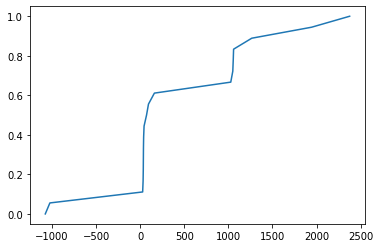
Now if an image is something like a bimodal distribution then it should have values like:
(this is an example)
[-200,-199,-199,-199,-198,-198,-198,-198,-198,-197,-196,-20,0,4,10,10,11,11,12,12,14,15]
and if the positions were like [2,4,6,8,10]
Then the bins become [-199, -199, -198, -196,0...]
Thus we can see how this function is designed to split values into groups, such that each group has around the same number of values.
Always plenty to learn from fastai