Hi,
I have a working prototype for possible SparseDataset and LinearSparse for creating dataset (mostly for tabular) from scipy.sparse using a custom sparse_collate_fn , would this be something desired or worth adding as a feature ?
I read a lot of issues in Pytorch forums and came up with the ideas above for handling sparse datasets.
Some use cases, why would someone want sparse dataset when we have embeddings ?:
1) Extracting features from CNN models like ResNet, if you have n different models it will make n*2048 sized vector as input which has lot of zeros since it’s after relu.
2) Text features which are non-sequential and variable length. Here we can use BOW-TFIIDF, of course you can work your way around with more sophisticated models like LSTMs or Transformers but BOW-TFIDF is most of the will give you a strong and fast baseline.
3 Will think more…
Why custom collate_fn?:
Because default_collate will fail. The way I do is i store scipy.sparse data inside dataset:
def sparse_collate_fn(dataset):
"""
dataset: [self.dataset[i] for i in indices]
"""
x,y = list(zip(*dataset))
sparse_stacked = scipy.sparse.vstack(x)
torch_sparse_stacked = torch_from_scipysparse(sparse_stacked, size=sparse_stacked.shape, device=0)
torch_y = torch.FloatTensor(np.concatenate([y]))
return torch_sparse_stacked, torch_y
def torch_from_scipysparse(sparse_matrix, *args, **kwargs):
"""sparse_matrix: scipy sparse matrix """
sparse_matrix = sparse_matrix.tocoo(copy=False)
row,col,values = sparse_matrix.row, sparse_matrix.col, sparse_matrix.data
i = torch.LongTensor(np.vstack([row, col]))
v = torch.FloatTensor(values)
return torch.sparse_coo_tensor(i, v, *args, **kwargs)
Why custom LinearSparse?:
Because at the moment autograd supports some ops only, which can be find here: https://github.com/pytorch/pytorch/issues/9674. This is an open issue and i bet in near future they will support all modules. But if you create nn.Linear layer move it to say device:0 and do backward() will fail. The reason is, at least what I remember from forums, is that when you move variables intermediate variables are created and backward fails ? (maybe someone can explain further). So LinearSparse will allow us to move the module to GPU as we create it, it uses torch.mm and pretty similar to nn.Linear:
class LinearSparse(nn.Module):
def __init__(self, in_features, out_features, bias=True, **kwagrs):
"""allow args and kwargs for weight and bias construction"""
super(LinearSparse, self).__init__()
self.in_features = in_features
self.out_features = out_features
self.weight = Parameter(torch.rand(in_features, out_features, **kwagrs))
if bias:
self.bias = Parameter(torch.rand(out_features, **kwagrs))
else:
self.register_parameter('bias', None)
self.reset_parameters()
def reset_parameters(self):
stdv = 1. / math.sqrt(self.weight.size(1))
self.weight.data.uniform_(-stdv, stdv)
if self.bias is not None:
self.bias.data.uniform_(-stdv, stdv)
def forward(self, input):
"""input: Sparse (S), weight: Dense(S)"""
return torch.mm(input, self.weight) + self.bias
def extra_repr(self):
return 'in_features={}, out_features={}, bias={}'.format(
self.in_features, self.out_features, self.bias is not None
)


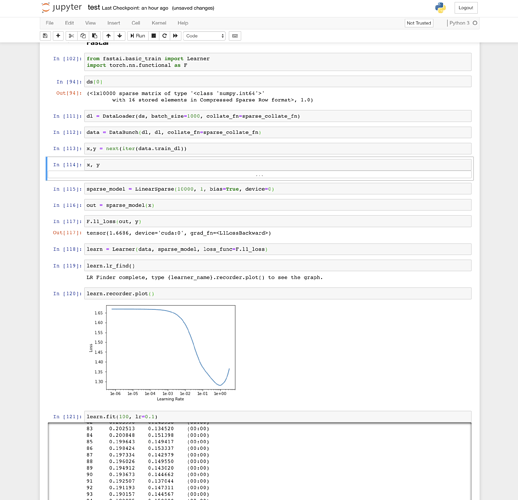


 So my point was introduce this behaviour into the library to make interoperation with “standard” PyTorch classes more simple. Like, to make it possible drop-in replacement of fastai loaders and datasets (maybe models also?) with plain PyTorch objects.
So my point was introduce this behaviour into the library to make interoperation with “standard” PyTorch classes more simple. Like, to make it possible drop-in replacement of fastai loaders and datasets (maybe models also?) with plain PyTorch objects.