Here is link to repo… One could improve progress bar… Hope it helps!
the function at the end creates following graph (using standard wd or provided by users)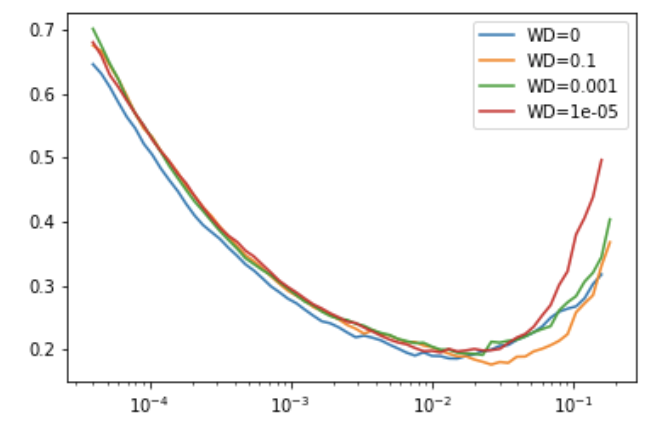
Here is link to repo… One could improve progress bar… Hope it helps!
the function at the end creates following graph (using standard wd or provided by users)