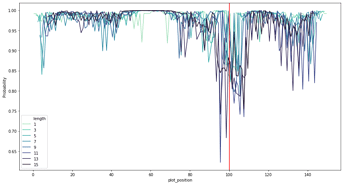
I’ve done ULMFiT on genomic sequences (link) which has worked well for classification of sequences up to a few kb. I was also able to infer important sequence regions by perturbing regions of the sequence to see the impact on the classification confidence. Here’s an example for promoter classification, where the red line shows the true transcription start site.
How much sequence information are you dealing with? I’ve been curious about the ability of LSTM models for genomics classification to scale to extremely long sequences, given the difficulty of maintaining information over a large number of time steps.
You might also consider looking into how SVD can be used on genomics data to reduce dimensionality and extract features in an unsupervised fashion.