Worked perfectly! Thank you!
Just me. Based in SF.
I’m trying to extend the deeplearning1 notebook mnist.ipynb and am running into some bugs. On the line
lm.fit_generator(batches, batches.N, nb_epoch=1, validation_data=test_batches, nb_val_samples=test_batches.N)
I first had the error message
AttributeError: 'NumpyArrayIterator' object has no attribute 'N'
which is raised on this thread, with a solution but not an explanation. I changed .N to .n as suggested there and then had an exception on the same line:
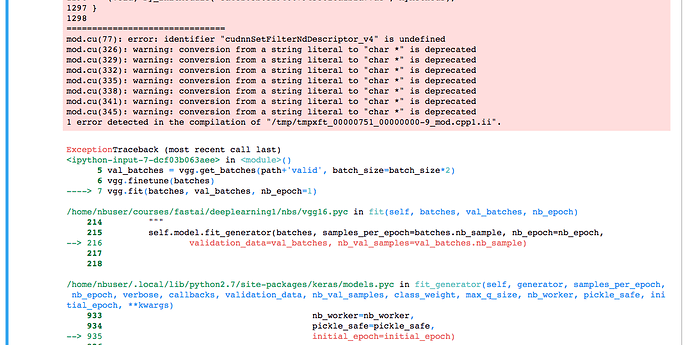
error: identifier "cudnnSetFilterNdDescriptor_v4" is undefined
which seems to be a problem with incompatibility between different versions of the libraries, discussed on github.
I’ve already been warned:
Using gpu device 0: Tesla K80 (CNMeM is enabled with initial size: 95.0% of memory, cuDNN 6021)
/home/nbuser/.local/lib/python2.7/site-packages/theano/sandbox/cuda/__init__.py:631: UserWarning:
Your cuDNN version is more recent than the one Theano officially supports.
If you see any problems, try updating Theano or downgrading cuDNN to version 5.1.
warnings.warn(warn)
I’m wary of fooling around trying to change library versions when I don’t fully understand the crestle environment, and I don’t know the command to issue to downgrade cuDNN or to upgrade theano. @anurag, can you suggest a good way to work around this? Thanks!
Hi @anurag. Thanks for setting up this service, so far it seems great. I’ve run into one problem however.
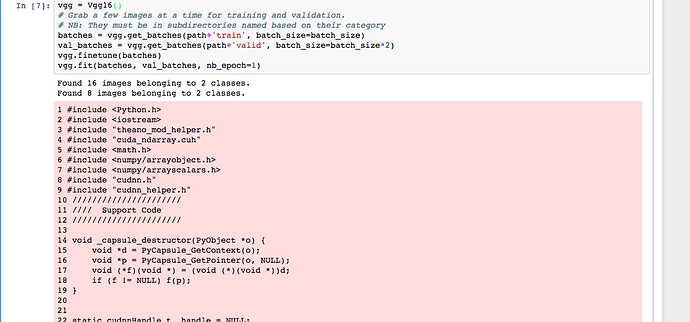
I’m getting this error when running input 7 on lesson 1 notebook:
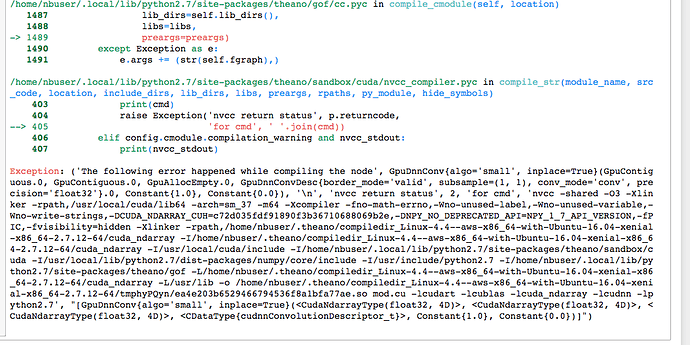
between the pictures posted, the notebook scrolls through the various errors in each line of the script, I but I’ve just shown the first and last extremes of this.
Any idea what might be going on?
Thanks!
I’m on crestle and I’m getting the same error. It’s so far down in the libraries I can’t guess what the problem really is. Any help would be appreciated.
Thanks for the reports. I should have a fix out shortly.
Should be fixed if you restart Jupyter.
Thanks Anurag – appreciate the quick attention!
Hey @anurag first of all thank you very much for your contribution. Currently doing Lesson 1. I would like to know how or where are we reading the files on? Because we define path = “/datasets/fast.ai/dogscats/” and I can’t seem to find that directory in the files… So where should I store files if later on I would like to run code with my own dataset?
Thanks in advance.
Thank you so much!
Just out of curiosity, what was causing the problem?
Yes, that’s fixed it for me, too, although I still have to change “.N” to “.n”.
Thanks!
Emrys
Whoops, I replied to @taguilera when I should have been replying to @anurag. Thanks, @anurag!
@taguilera, /datasets/fast.ai/dogscats/ is at the whole filesystem root, as opposed to the per-user root that you see with the Jupyter file browser. If you hit new terminal in Jupyter, you get a linux shell. Type
ls /
to see the filesystem root and you should see datasets as one of the directories there, and you can cd around to explore. Type
ls ~
to see your own user’s files, which is what the Jupyter file browser displays.
If you store your own data anywhere in your own user’s files, it stays stored there. At least, mine does.
Good Luck!
Emrys
Hi Anurag,
I have just started using Crestle…First of all I would like to thank you for making this. I find it very helpful. Just some doubt, is there any way to access file from local drives…
Thnx…
I would transfer data to a public host and use wget to download it in the terminal.
I upgraded to cuDNN 6, which Theano doesn’t like. Downgraded back to v5. This should all be fixed in the next version of Part 1 which does not use Theano (which has ceased active development anyway).
Ahhh I see.
thnx a lot…
anyone know how i can get kaggle data into this? i’m trying the Statefarm distracted driver competition and the data is 4gb. https://www.kaggle.com/c/state-farm-distracted-driver-detection/data
Edit: Just noticed that the Statefarm data is already there in the kaggle datasets folder - awesome! Only problem is it seems to be in zipped folders and I can’t seem to find the train and validation sets.
Thanks again for everything you’ve done. Truly couldn’t be more grateful.
There was an issue with kaggle’s statefarm data earlier. It’s fixed and everything is unzipped under /datasets/kaggle/state-farm-distracted-driver-detection. Hope this helps!