Hi there,
I’m quite new to FastAI and I am hoping to use a model to classify fluorescent images of cells. Essentially for any given cell, there are 4 fluorescent channels (One each for the nucleus, actin filaments, golgi body and mitochondria). What I would like to do is train a model on each fluorescent channel separately and that then combines into one output function.
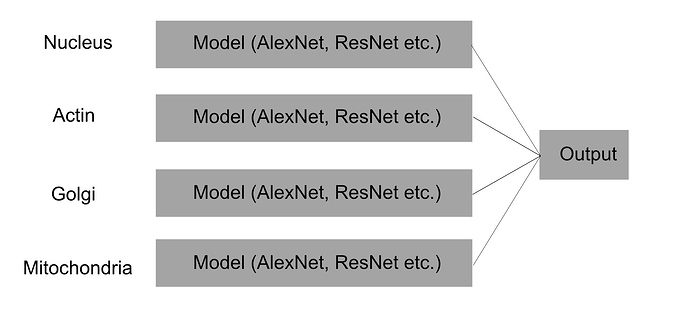
Essentially I want a create a large model that can look at the nucleus, actin filaments, golgi body and mitochondria with a separate model for each, independent of one another until the final classification layer (I hope I have explained that appropriately, I have attached an image for some visual understanding)
Note: Each nucleus channel image for example, is exactly the same size as the corresponding actin, golgi and mitochondria image for a given cell.
If anyone could point me in the right direction or requires more information, please let me know.
Cheers.