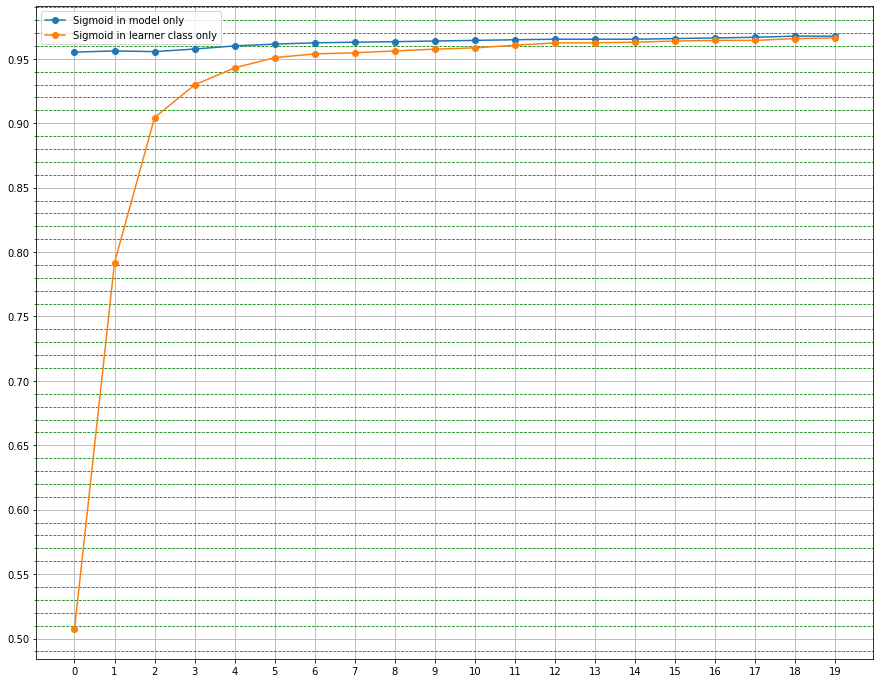
I was trying to build the learner from scratch, but there seems to be something weird happening. Below is the Code for my Leaner and I have also included the other necessary information.
class Learner_try:
def __init__(self, dl, model, opt):
self.dl_train = dl[0]
self.dl_valid = dl[1]
self.model = model
self.opt = opt(self.model.parameters(), lr = 0.1)
def mnist_loss(self, preds, targets):
preds = preds.sigmoid()
return torch.where(targets==1, 1 - preds, preds).mean()
def batch_accuracy(self, x, y):
preds = x.sigmoid()
correct = (preds>0.5) == y
return correct.float().mean()
def validate_epoch(self):
accs = [self.batch_accuracy(self.model(x), y) for x,y in self.dl_valid]
return round(torch.stack(accs).mean().item(), 4)
def cal_grad(self, x, y):
preds = self.model(x)
loss = self.mnist_loss(preds, y)
loss.backward()
def train_epoch(self):
for x, y in self.dl_train:
self.cal_grad(x, y)
self.opt.step()
#self.opt.zero_grad() #This is the step which is acting wierd
def fit(self, epochs):
for i in range(epochs):
self.train_epoch()
print(self.validate_epoch(), end = " ")
simple_net = nn.Sequential(nn.Linear(28 * 28, 30),
nn.ReLU(),
nn.Linear(30, 1),
nn.Sigmoid())
opt = SGD
learn = Learner_try(dls, simple_net, opt = opt)
learn.fit(20)
I have used the SGD directly as my optimizer, when I fit the learner without the “self.opt.zero_grad” step in the “train_epoch” method it works fine(getting me a score above 0.96 - 0.97- which it should actually do) but when I run it with the “self.opt.zero_grad” step it kind of sticks at one point getting a value eg:0.4957 for “n” number of epochs.
I might have done something wrong or missed something, First I thought that SGD must be handling step() and zero_grad() together but when I checked the source code SGD when it returns to Optimizer - Optimizer does have the zero_grad() method.
Any help here would be appreciated.