Hi,
in one of my recent projects I’m trying to build a autoencoder to encode long gene sequences in a smaller latent vector to search sequences and try to find similarities between different sequences. Since the Sequences are quite long (up to >60.000) I used a combination of conv layers and LSTM units. For a proof of concept I limited the data to sequences with a length of <= 3000. The dataset I used can be found here.
As loss function I’m using cross_entropy and the loss is decreasing within the first epochs, however I cant get it any lower than 0.6 on train and validation data. And the reconstruction from the latent vector does not look any close to the input:
print(val_df.seq.values[0])
ATGTGTCCCCGAGCCGCGCGGGCGCCCGCGACGCTACTCCTCGCCCTGGGCGCGGTGCTGTGGCCTGCGGCTGGCGCCTGGGAGCTTACGATTTTGCACACCAACGACGTGCACAGCCGGCTGGAGCAGACCAGCGAGGACTCCAGCAAGTGCGTCAACGCCAGCCGCTGCATGGGTGGCGTGGCTCGGCTCTTCACCAAGGTTCAGCAGATCCGCCGCGCCGAACCCAACGTGCTGCTGCTGGACGCCGGCGACCAGTACCAGGGCACTATCTGGTTCACCGTGTACAAGGGCGCCGAGGTGGCGCACTTCATGAACGCCCTGCGCTACGATGCCATGGCACTGGGAAATCATGAATTTGATAATGGTGTGGAAGGACTGATCGAGCCACTCCTCAAAGAGGCCAAATTTCCAATTCTGAGTGCAAACATTAAAGCAAAGGGGCCACTAGCATCTCAAATATCAGGACTTTATTTGCCATATAAAGTTCTTCCTGTTGGTGATGAAGTTGTGGGAATCGTTGGATACACTTCCAAAGAAACCCCTTTTCTCTCAAATCCAGGGACAAATTTAGTGTTTGAAGATGAAATCACTGCATTACAACCTGAAGTAGATAAGTTAAAAACTCTAAATGTGAACAAAATTATTGCACTGGGACATTCGGGTTTTGAAATGGATAAACTCATCGCTCAGAAAGTGAGGGGTGTGGACGTCGTGGTGGGAGGACACTCCAACAC…
latent = ae.encode(x=val_df.ids.values[0])
ae.sample(latent)
ATGGAGGGGGGGGGGGGGGGGGGGGGGGGCCTGGCCGGGGGGGGGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCTGGCCCTGGCCCTGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAAGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAGGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAAAAAAAAAAAAAAAAGAAGAAGAAAAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAAAAGAAAAAAAAAAAAAAAAAAAAAAAAAAAAAGAAGAAGAAGAAGAAGAAGAAAAAAAAAAAGAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAGAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAGAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAG…
Are there any suggestions on improving the encoding? This is the model I’m currently using:
class AutoEncoder(nn.Module):
def __init__(self,vocab_size, embed_size, latent_size,filters,decoder_out_filters,
seqLength,pad_idx,pre_pad,
stoi,itos,GO_TOKEN,STOP_TOKEN):
super().__init__()
self.latent_size = latent_size
self.vocab_size = vocab_size
self.embed_size = embed_size
self.seqLength = seqLength
self.pad_idx = pad_idx
self.pre_pad = pre_pad
self.itos = itos
self.stoi = stoi
self.GO = GO_TOKEN
self.STOP = STOP_TOKEN
self.decoder_out_filters = decoder_out_filters
self.encFilters = filters.copy()
filters.reverse()
self.decFilters = filters.copy()
self.embed = nn.Embedding(self.vocab_size, self.embed_size)
self.embed.weight = xavier_normal_(self.embed.weight)
self.encoder = Encoder(embed_size,latent_size,self.encFilters)
self.decoder = Decoder(vocab_size,latent_size,50,2,embed_size,self.decFilters,out_filters=decoder_out_filters)
def forward(self,encIn,decIn):
encIn = self.embed(encIn)
encOut = self.encoder(encIn)
decIn = self.embed(decIn)
decOut = self.decoder(encOut,decIn)
return decOut
def encode(self,x):
if isinstance(x,list):
x = np.asarray(x)[None]
res = np.zeros((len(x), self.seqLength), dtype=x[0].dtype) + self.pad_idx
for i,o in enumerate(x):
if self.pre_pad: res[i, -len(o):] = o
else: res[i, :len(o)] = o
emb = self.embed(V(T(res)))
latent = self.encoder(emb)
return latent
def sample(self,latent):
final_state = None
x = self.embed(V(T([self.stoi[self.GO]])[None]))
result = [self.GO]
cnn_out = self.decoder.conv_decoder(latent)
for var in t.transpose(cnn_out,0,1):
out, final_state = self.decoder.rnn_decoder(var.unsqueeze(1),decoder_input=x,initial_state=final_state)
out = F.softmax(out.squeeze())
out = to_np(out)
idx = out.argmax()
char = self.itos[idx]
result.append(char)
if char == self.STOP:
break
x = self.embed(V(T([idx])[None]))
return ''.join(result)
Encoder:
class ConvBlock(nn.Module):
def __init__(self,prev,nf,kernelsize,stride):
super().__init__()
self.convblock = nn.Sequential(nn.Conv1d(prev, nf, kernelsize, stride),
nn.BatchNorm1d(nf),
nn.ELU())
def forward(self,x):
return self.convblock(x)
class Encoder(nn.Module):
def __init__(self, embed_size, latent_size,filters):
super().__init__()
self.embed_size = embed_size
self.latent_size = latent_size
filters = [self.embed_size]+filters+[self.latent_size]
convblocks = []
for i in range(1,len(filters)):
if i == 9:
kernel = 2
else:
kernel = 4
convblocks.append(ConvBlock(filters[i-1],filters[i],kernel,2))
self.cnn = nn.Sequential(*convblocks)
def forward(self,x):
x = t.transpose(x, 1, 2)
result = self.cnn(x)
result = result.squeeze(2)
return result
Decoder:
class DeconvBlock(nn.Module):
def __init__(self,prev,nf,kernelsize,stride,padding):
super().__init__()
self.convblock = nn.Sequential(nn.ConvTranspose1d(prev, nf, kernelsize, stride,0, padding),
nn.BatchNorm1d(nf),
nn.ELU())
def forward(self,x):
return self.convblock(x)
class Decoder(nn.Module):
def __init__(self,vocab_size, latent_variable_size, rnn_size, rnn_num_layers, embed_size, filters,out_filters):
super().__init__()
self.vocab_size = vocab_size
self.latent_variable_size = latent_variable_size
self.rnn_size = rnn_size
self.embed_size = embed_size
self.rnn_num_layers = rnn_num_layers
self.out_filters = out_filters
filters = [self.latent_variable_size]+filters+[self.out_filters]#+[self.vocab_size]
convblocks = []
for i in range(1,len(filters)):
kernel = 4
padding = 0
if i == 2:
kernel = 2
if i in [2,3,5,6,7,9]:
padding = 1
convblocks.append(DeconvBlock(filters[i-1],filters[i],kernel,2,padding))
self.cnn = nn.Sequential(*convblocks)
self.rnn = nn.GRU(input_size=self.out_filters + self.embed_size,
hidden_size=self.rnn_size,
num_layers=self.rnn_num_layers,
batch_first=True)
self.hidden_to_vocab = nn.Linear(self.rnn_size, self.vocab_size)
def forward(self,latent_variable,decoder_input):
aux_logits = self.conv_decoder(latent_variable)
logits, _ = self.rnn_decoder(aux_logits, decoder_input, initial_state=None)
return logits
def rnn_decoder(self, cnn_out, decoder_input, initial_state=None):
logits, final_state = self.rnn(t.cat([cnn_out, decoder_input], 2), initial_state)
[batch_size, seq_len, _] = logits.size()
logits = logits.contiguous().view(-1, self.rnn_size)
logits = self.hidden_to_vocab(logits)
logits = logits.view(batch_size, seq_len, self.vocab_size)
return logits, final_state
def conv_decoder(self, latent_variable):
latent_variable = latent_variable.unsqueeze(2)
out = self.cnn(latent_variable)
out = t.transpose(out, 1, 2).contiguous()
return out
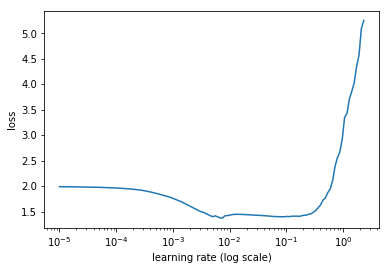
LR-Find:
I tried several parameter settings, which all led to a loss around 0.6-ish.
layers = [128,128,128,256,256,256,512,512,512]
ae = AutoEncoder(len(itos),200,512,layers,
decoder_out_filters = 512,
seqLength=3000,
pad_idx=0,
pre_pad=False,
GO_TOKEN=GO,
STOP_TOKEN=STOP,
itos=itos,
stoi=stoi)
learn = RNN_Learner(md, SingleModel(to_gpu(ae)))
learn.crit = celoss
learn.opt_fn = optim.Adam
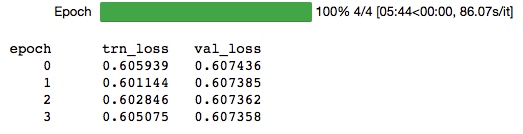
lr = 1e-3
learn.fit(lr,1,cycle_len=3,cycle_mult=3,use_wd_sched=False,use_clr=(20,10))
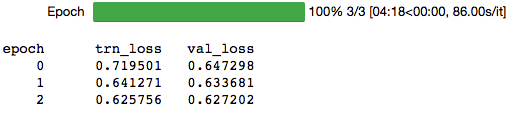
lr = 1e-4
learn.fit(lr,1,cycle_len=4,cycle_mult=2)
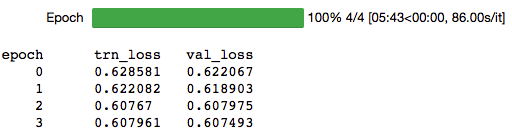
lr = 1e-6
learn.fit(lr,1,cycle_len=4,cycle_mult=2)