I don’t have an exact plan as of now. (Also I’m home for holidays and my family does not appreciate me spending time on my laptop while I’m here so it’s hard to really get started on anything)
Although I do have at least one very specific thing in mind. And I think it’s pretty cool.
So, I wanted to test out a modification to BatchNorm layers after doing Part-2 of the course. I was looking for a function that can somehow scale all it’s input to between -1 and 1 without any parameters. Naturally, I turned to activation functions such as Sigmoid, exp(-x^2), Tanh, etc. After some experimentation in the 07_batchnorm notebook itself, what finally seemed to work reliably was applying Tanh followed by the usual scaling and shifting with mults and adds in BatchNorm (Essentially, I replaced the normalization part of a BatchNorm layer with a Tanh). The final results looked pretty (very) close to the usual BatchNorm at least for the network in that Notebook
97.35% vs 97.66% accuracy for 5 epochs each.
Since this looked encouraging, I wanted to try this modification for all the usual networks (Resnets, Inception, etc.) and see if this generalizes with datasets that are more serious than MNIST.
But I didn’t want to have to write new model definitions for all these architectures as I figured that I’ll be needing to write a whole lot more definitions this way and that will slow me down.
So I went ahead and just tried setting the bn* modules in the factory PyTorch resnet modules to a Tanh layer.
THIS DID NOT WORK.
It seemed to break some of the skip-connection logic. I realized that the same issue will probably crop up in other networks too.
After a bit of fiddling around, I found this bit of code to work:
def recursive_getattr(obj:nn.Module, name:str):
""" getattr for nested attributes with `.` in their names """
sequence = name.split('.')
if len(sequence) == 0 or name == '': return obj
for attr in sequence: obj = getattr(obj, attr)
return obj
def recursive_setattr(obj:nn.Module, name:str, new_attr):
""" setattr for nested attributes with `.` in their names """
sequence = name.split('.')
obj = recursive_getattr(obj, '.'.join(sequence[:-1]))
setattr(obj, sequence[-1], new_attr)
def modify_network(m:nn.Module, replace_func:Callable, condition:Callable=None)->nn.Module:
""" modifies `m` by replacing each module that satisfies `condition`
with replace_func(module) """
if condition is None:
# If `replace_func` has a condition in-built, it should return
# the passed module as it is. This is to prevent a recursion issue.
condition = (lambda x: not x == replace_func(x))
modules = list(m.named_modules())
if len(modules) == 1:
return replace_func(m)
for name, module in modules:
if condition(module):
recursive_setattr(m, name, replace_func(module))
return m
After this I can do what I originally intended with very little code:
def bn_to_tanh(m):
if isinstance(m, nn.BatchNorm2d):
nf = m.num_features
m = TanHNorm(nf) # A module I defined elsewhere
return m
m = models.resnet18(pretrained=False)
modify_network(m, bn_to_tanh)
This gives me back a working resnet18 with all the BatchNorm layers replaced with a suitable tanh layer. I have yet to test it with other kinds of architectures.
In essence, this allows me to take models created by anyone with whatever logic they might have used in their script, and then just modify them as needed with very little code. This can be handy when trying out new activation functions, particular modifications to cnn layers or just in general adding/modifying arbitrary parts of existing networks with little effort. (Can this be added to Fastai?)
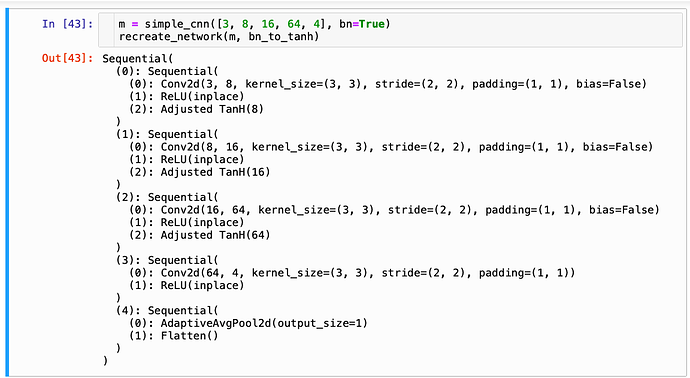
It does work with FastAI’s (V1) simple_cnn function:
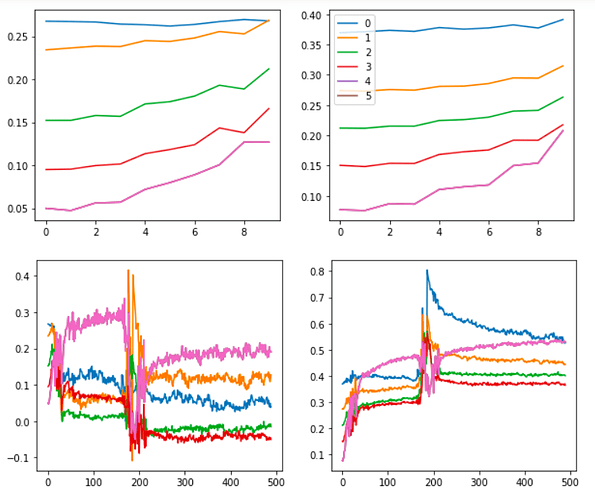
Also, the graphs for the tanh experiment look like this:
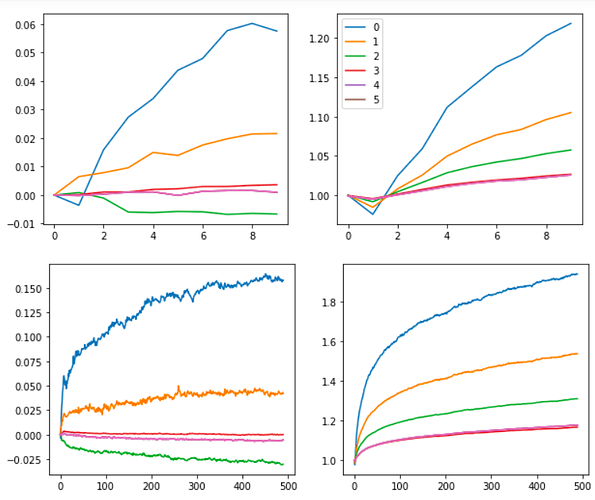
As opposed to the usual
BatchNorm:
It’s not as smooth, but I think this is worth exploring further. The core advantage of using this approach is that there are no variance or mean parameters or calculations (since that’s generally what causes a lot of problems with batch sizes and generalization). By getting rid of those parameters, we may get rid of those problems too. (I’m still learning though. Please, please correct me if this sounds naive or wrong)
I want to start running experiments with resnets on Imagewoof and Imagenette using this approach as soon as possible but right now I’m all caught up in preparing to go back to college. Hopefully, I’ll have something tangible to show for soon enough.
As far as writing the blog is concerned, I was hoping to discover new things as I go along ripping apart some of the core functions in nn.Module just like I did with matplotlib. A significant part of that was covered in Part-2 of the course itself, so that might help here too.
(I’m thinking of exploring other libraries as well - Jax: Numpy with Autograd, XLA and GPUs, streamlit: easily creating data science GUI tools/apps)

 .
.