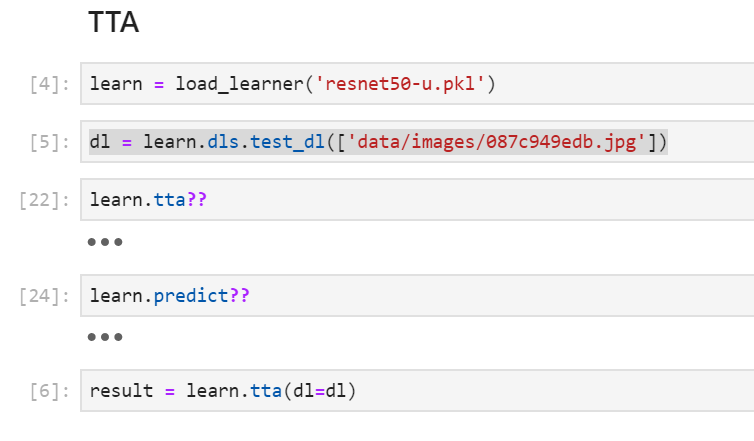
You should make a test_dl then do learn.tta(dl=dl)
Can you point me to an example of how to create a test_dl out of a simple image path?
If you want it for one image (rather than many), wrap your path in an array as you pass it in:
dl = learn.dls.test_dl([fname])
Re:example, it’s right in predict’s source code:
learn = load_learner('model.pkl')
dl = learn.dls.test_dl([filepath])
predictions = learn.tta(dl=dl, n=4)
Mmm… works fine in the notebook but throws an error in the script
epoch train_loss valid_loss f1_score time
Traceback (most recent call last):
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 182, in _do_epoch_validate
self.dl = dl; self('begin_validate')
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 134, in __call__
def __call__(self, event_name): L(event_name).map(self._call_one)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 377, in map
return self._new(map(g, self))
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 327, in _new
def _new(self, items, *args, **kwargs): return type(self)(items, *args, use_list=None, **kwargs)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 47, in __call__
res = super().__call__(*((x,) + args), **kwargs)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 318, in __init__
items = list(items) if use_list else _listify(items)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 254, in _listify
if is_iter(o): return list(o)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 220, in __call__
return self.fn(*fargs, **kwargs)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 137, in _call_one
[cb(event_name) for cb in sort_by_run(self.cbs)]
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 137, in <listcomp>
[cb(event_name) for cb in sort_by_run(self.cbs)]
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/callback/core.py", line 24, in __call__
if self.run and _run: getattr(self, event_name, noop)()
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/callback/progress.py", line 26, in begin_validate
def begin_validate(self): self._launch_pbar()
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/callback/progress.py", line 34, in _launch_pbar
self.pbar = progress_bar(self.dl, parent=getattr(self, 'mbar', None), leave=False)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastprogress/fastprogress.py", line 226, in __init__
super().__init__(gen, total, display, leave, parent, master)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastprogress/fastprogress.py", line 24, in __init__
parent.add_child(self)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastprogress/fastprogress.py", line 264, in add_child
self.child.prefix = f'Epoch {self.main_bar.last_v+1}/{self.main_bar.total} :'
TypeError: unsupported operand type(s) for +: 'NoneType' and 'int'
During handling of the above exception, another exception occurred:
Traceback (most recent call last):
File "run.py", line 118, in <module>
run()
File "run.py", line 84, in run
predictions = learn.tta(dl=dl, n=4)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 542, in tta
with dl.dataset.set_split_idx(1): preds,targs = self.get_preds(dl=dl, inner=True)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 229, in get_preds
self._do_epoch_validate(dl=dl)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 185, in _do_epoch_validate
finally: self('after_validate')
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 134, in __call__
def __call__(self, event_name): L(event_name).map(self._call_one)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 377, in map
return self._new(map(g, self))
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 327, in _new
def _new(self, items, *args, **kwargs): return type(self)(items, *args, use_list=None, **kwargs)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 47, in __call__
res = super().__call__(*((x,) + args), **kwargs)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 318, in __init__
items = list(items) if use_list else _listify(items)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 254, in _listify
if is_iter(o): return list(o)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 220, in __call__
return self.fn(*fargs, **kwargs)
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 137, in _call_one
[cb(event_name) for cb in sort_by_run(self.cbs)]
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/learner.py", line 137, in <listcomp>
[cb(event_name) for cb in sort_by_run(self.cbs)]
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/callback/core.py", line 24, in __call__
if self.run and _run: getattr(self, event_name, noop)()
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastai2/callback/core.py", line 96, in after_validate
if not self.save_preds: self.preds = detuplify(to_concat(self.preds, dim=self.concat_dim))
File "/home/ec2-user/anaconda3/envs/snakes/lib/python3.6/site-packages/fastcore/foundation.py", line 234, in __getattr__
if attr is not None: return getattr(attr,k)
AttributeError: 'Learner' object has no attribute 'preds'Question: in the notebook is the learn the same one as trained? (IE you did not do load_learner)
What do you mean?  I always do learn = load_learner(‘model.pkl’)
I always do learn = load_learner(‘model.pkl’)
- It is curious because I see in the terminal 4 times the prediction bar running (tta with n=4) but afterwards the error is thrown
Hmmm… not sure there then. May be a bug in the library? Signaling @sgugger if he’s available, else perhaps I should start pinging Jeremy 
Hi!
I was following the notebook on Object Detection in class 6 and there defines the data as follows:
pascal = DataBlock(blocks=(ImageBlock, BBoxBlock, BBoxLblBlock),
splitter=RandomSplitter(),
get_items=get_train_imgs,
getters=getters,
item_tfms=item_tfms,
batch_tfms=batch_tfms,
n_inp=1)
dls = pascal.dataloaders(path/'train')
Instead, I would like to do the split using ColSplitter(), since the DataFrame I am using is as follows:
image labels x0 y0 x1 y1 is_valid
0 Image1194.png A4 499 452 546 303 True
1 Image0847.png A6 1075 429 243 49 False
2 Image1071.png A4 642 864 147 98 False
3 Image1071.png A4 636 643 153 104 False
4 Image1195.png A4 581 127 286 155 False
... ... ... ... ... ... ... ...
But I don’t know how to integrate this type of split into the current format of the bounding boxes. Since when creating the dataloaders in this case I need both, the DataFrame and the path.
How can I integrate these two things at the same time? Thanks!
I’ve been going through the Vision fastai v2, but I don’t see any support for using pathnames and labels from a csv (pandas dataframe)? Has this been removed in v2?
No, this is now ColReader. There are three example in the multilabel notebook here
Thanks, the ColReader was what I was looking for.
I’m still extremely confused as to how to use learn.validate.
Sorry for this longer post. My tldr is figuring out how to pass a new dataloader and get learn.validate metrics out from that.
In fastai v1, I could do this.
learn.validate(train_data.train_dl) learn.validate(train_data.valid_dl) learn.validate(test_data.train_dl)
Where I can rerun learn.validate to get the final metrics for my training and validation set, as well as on another test dataset. I don’t use tst_dl because it needs to be defined when constructing the dataloaders, so I create another ‘test_data’ dataloader and assign the test dataset to be the train_dl object
Now, in fastai v2, the following gives me a “TypeError: object of type ‘DataLoaders’ has no len()” error
learn.validate(dl=train_dls)
where train_dls is the object
pets = DataBlock(blocks=(ImageBlock, CategoryBlock),
get_x=ColReader("file"),
get_y=ColReader("category"),
splitter=RandomSplitter(),
item_tfms=item_tfms,
batch_tfms=batch_tfms)
train_dls = pets.dataloaders(train_df, bs=bs)
This is super weird as train_dls is the exact same dataloader I passed into learn in order to train it! The following code works:
learn.validate(ds_idx=0)
learn.validate(ds_idx=1)
Except that this only reruns the metrics for the training and validation dataset, with no option to inject/replace a new dataloader (or even the same one). I tried searching for instances where learn.validate was used and found this in the notebooks:
dl = learn.dls.test_dl(df2) learn.validate(dl=dl)
Except for my case I do not have a learn.dls.test_dl object, or even learn.dls.train_dl. All I have is learn.dls.train and learn.dls.train_ds.
The DL wants a single DataLoader. Not a set of DataLoaders (what you did in the first example). You could have done dl=train_dls[0] for instance for the training data
For the test, learn.dls.test_dl generates a test DataLoader for you to use, passing it into learn.validate(dl=dl). IIRC you need to pass is_labeled=True to do so.
Train_dl and valid_dl are no longer used. They are .train and .valid
Kind of confused as to why there is 2 types of resizing methods,for example:
batch_tfms = [*aug_transforms(size=224), Normalize.from_stats(*imagenet_stats)]
item_tfms = RandomResizedCrop(460, min_scale=0.75, ratio=(1.,1.))
From the input going into the network, I’m pretty sure the final size after data augmentation is 224x224. So why are we doing another RandomResizedCrop?
I assume the sequence is original size -> RandomResizedCrop to 460x460 -> Resize to 224x224 -> Feed into model?
Correct. It’s a method called Pre-Sizing. Jeremy covers it in fastbook/course-v4
Assuming you are using CUDA (GPU), one big difference is that item_tfms happens on CPU, item-by-item (where each item might have different initial sizes), whereas batch_tfms, with the items now already ‘pre-sized’ to the same size as @muellerzr mentioned, can go onto the GPU as a batch and get resized (to the final 224 in your sample code) more efficiently.
Yijin
I’ve been looking through batchnorm, and could not find any robust guidance of batchnorm issues at test time. If you’ve come across better guidance, do let me know. Thanks!
First, kudos to Jeremy who looked into batchnorm issues during transfer learning. You can imagine that if you don’t re-learn the batchnorm layers during transfer learning that will cause issues especially if the original dataset and the new dataset are from different distribution. In his fastai v2 paper: “One area that we have found particularly sensitive in transfer learning is the handling of batch-normalization layers [3]. We tried a wide variety of approaches to training and updating the
moving average statistics of those layers, and different configurations could often change the error rate by as much as 300%. There was only one approach that consistently worked well across all datasets that we tried, which is to never freeze batch-normalization layers, and never turn off the updating of their moving average statistics”
Here’s my biggest issue with batchnorm being used everywhere. Even with Jeremy’s fix that helps training, nobody is talking about inference. Yes, at test time you use the stats observed during training and apply it, but that ONLY works if your test data is within the same if not identical distribution as your training data. One solution I see is to precompute the stats on the test data and use that during inference… but how? Even if I could do it, the model was trained on the running stats of the train data, so this incompatibility might lead to other issues.
The fast.ai leaderboards are also illustrating this point perfectly. We have no evaluations on how well models generalize to slightly different distribution test data. Test data of ImageNette and ImageWoof all belong to the same distribution.
It’s also frustrating at many established pre-trained model comes with batchnorm layers. 
If anyone wants to help out some, here is that Gaussian keypoint implementation I mentioned (realized today I never made the thing public). Folks who are familiar with pose detection I could certainly use your help. The implementation is based on HRNet:
I’m curious if this is a bug? I’m running some inference preds, where I remove all data augmentation during test time and getting drastically different results when I use:
batch_tfms = [*aug_transforms(mult=0.0, size=224,
do_flip=False, flip_vert=False, max_rotate=0.,
min_zoom=1.0, max_zoom=1.0,
max_lighting=0., max_warp=0.,
p_affine=0., p_lighting=0.,
pad_mode='reflection', min_scale=1.0),
Normalize.from_stats(*imagenet_stats)]
item_tfms = [Resize(540, method='squish')]
bs=64
Gives test pre/rec of 0.87/0.36
batch_tfms = [Normalize.from_stats(*imagenet_stats)]
item_tfms = [Resize(224, method='squish')]
bs=64
Gives test pre/rec of 0.81/0.85
I don’t understand why the first method gives wrong results? For the record val pre/rec is 0.84/0.82.
In the 1st method we resize to 540 then again to 224, but I disabled all data augmentations
In the 2nd method we resize directly to 224.
Shouldn’t both give the same results?
Ok, I found that this modification gives the correct results. Somehow resizing using aug_transforms in batch_tfms is not the same as using Resize in item_tfms?
batch_tfms = [*aug_transforms(mult=0.0, size=None,
do_flip=False, flip_vert=False, max_rotate=0.,
min_zoom=1.0, max_zoom=1.0,
max_lighting=0., max_warp=0.,
p_affine=0., p_lighting=0.,
pad_mode='reflection', min_scale=1.0),
Normalize.from_stats(*imagenet_stats)]
item_tfms = [Resize(224, method='squish')]
bs=64Looks good! Could this be used for key point regression where each item has a different number of key points?
That’s the goal. If you read the paper that is what’s done. The reason for the issue is when I train a model it has issues (training HRNet)