Here’s the link: https://youtu.be/-aCtDIgbxMw
Sorry guys! Laptop just pooped out on me. It’s a week. Coming back online momentarily 
Edit: and done, thanks for joining guys. @mgloria hopefully that answers your questions a bit better 
So happy to see you back!!  I had (once again) a fantastic time watching your video…!
I had (once again) a fantastic time watching your video…!
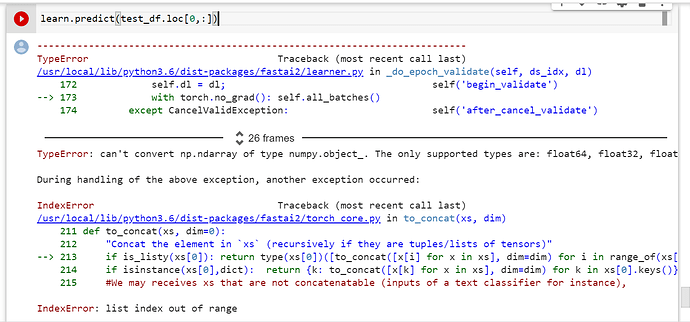
Does anybody know how to get predictions of a single row from the test_df (not several)? i.e. I would like to do something like this:
@mgloria try doing iloc IE df.iloc[100]
Interesting! I re-run today the notebook and both options worked!  I may have broken something yesterday while experimenting with the code.
I may have broken something yesterday while experimenting with the code.
Does anybody know what the outputs of learn.predict() are? I get from the documentation that it returns the following: full_dec,dec_preds,preds = learn.predict(test_df.iloc[100]) but I observe no difference between dec_preds and preds…
dec_preds is decoded from the loss function, and preds is the raw preds ![]()
What is the difference…? Not seeing your point 
The recording is available in this thread here (if you may not have access to this now that is okay). A basic summary is that when we call learn.get_preds (which predict does), we can pass in decoded=True. What this entails is that our loss function will decode the values. Why and how this could be different is say with CrossEntropy, we’ll actually run a softmax regularly, but our decoded values will also perform an argmax. Converting it to an actual class name is done through the dataloader’s decode_batch() function
Do check the contents of lesson 2 in form of Kaggle notebooks if interested
Next lecture will be on Wednesday night. We’re back and with a voice again so we’ll go to our regularly scheduled programming  (5pm CST)
(5pm CST)
I used Zach’s notebook to train a neural network on the adults dataset. I then extracted the embeddings to use as input for a random forest classifier to see if there is a difference in the accuracy vs a random forest without embeddings.
Is there a way to customize TabularPandas? I used the DataBlock to achieve the mapping between the categorical variables and the embeddings but the code I wrote feels quite clunky. Any tips appreciated 
@faib awesome work! (Reading through now) IIRC @Pak looked into this and found that it didn’t really make that much of a difference at the end of the day (back in v1) so the results aren’t too surprising.
Thank you!
I just realized I could write a processor like Categorify and append it to the procs argument in TabularPandas right?
I still have problems understanding where TabularPandas fits in. It’s not a high level API
like TabularDataLoaders nor does it belong to the DataBlock category. Does it logically sit below that or somewhere in between?
Somewhere in between. The role of TabularPandas is to prepare the data for being made into a DataLoader. It’s high-level API but it’s not a DataBlock (this is in development)
Got it! I’m looking forward to using the DataBlock API for tabular and being able to rely on using a unified interface 
When I try to add an additional metric to my learner, it results in below error during learn.fit():
*TypeError: unsupported operand type(s) for : ‘AccumMetric’ and ‘int’
Learn object defined as below:-
from fastai2.callback.all import *
learn = tabular_learner(dls,
layers=[1000,500],
metrics=[accuracy,RocAuc])
Need some help on this …
Hi Haroon!
I think this can help you:
I also get below error on passing the dropouts (ps, embed_p). I remember this working fine earlier.
TypeError: init() got an unexpected keyword argument 'ps’
learn = tabular_learner(dls,
layers=[1000,500],
ps=[0.001, 0.01],
embed_p=0.04,
metrics=[accuracy])
Thanks this fixed the issue.
I think the more recent version deprecates dropouts from the tabular_learner thus there will be no ps nor embed_p…